INTRODUCTION
Urinary tract infection is a common bacterial infection, known to affect various parts of the urinary tract and is observed in both genders. Even though both males and females are exposed to the infection, but women are more due to their morphology and reproductive physiology [1]. Besides one of the most prevalent infectious diseases found in outpatients is UTI. It is often caused by bacteria but may are also caused by fungi and viruses [2]. Gram-negative bacteria cause 90% of UTI cases while gram-positive bacteria cause only 10% of the cases. The frequency of the UTI depends on many factors that provide the prevalence of bacteria (for more than 10./ml) in the urine [3]. These bacteria cause UTI, and if not treated, the infection will spread and the patient will suffer severe consequences [4, 5, 6]. The most frequent isolated uropathogen is Escherichia coli, which accounts for 65-90% of urinary tract infections [7], followed by Staphylococcus species that constitute 10-15%. In addition, bacterial species such as Klebsiella, Pseudomonas, Proteus, and Enterococcus play a minor role in infection transmission [1]. UTI is linked to several factors, including age, parity, gravidity, pregnancy, and the presence of diseases that exacerbate the infection [1]. UTI antibiotic therapy is normally performed before the results of the microbiology examination are received. This medication, when used without a rational drug prescription, will result in antibiotic resistance and treatment failure [3, 8]. One of the best achievements in modern medicine was the discovery of antibiotics [9]. However, the availability and expanded use of antibiotics eventually contribute to microbial resistance to them. [9]. Urinary tract infection accounts for a large part of antibacterial drug consumption. Antibiotic resistance increases daily, according to irrational usage, particularly in countries with unrestricted antibiotic policies and no accurate plans [10] like Libya. Antimicrobial resistance is becoming a global danger to public health, according to the World Health Organization in 2014, and all countries have concentrated on this issue, which is a significant threat to modern medicine [11]. The excessive use of antibiotics is the first significant element in increasing microbial resistance. The other factor is wrong and unreasonably prescribed antibiotics. While UTI is a widespread disease, it is quickly cured by the reasonable use of antibiotics [4]. Identifying and analyzing UTI-causing bacteria is efficient in treating the resistance of antibiotics [12]. The purpose of this study was to determine the prevalence of bacteria associated with UTI cases and their antibiotic susceptibility pattern.
MATERIALS AND METHODS
STUDY SITTING
The study population was drawn from patients attending Tripoli Medical Center, the hospital in Tripoli city - Libya. Overall, 200 untreated outpatients with different clinical symptoms of UTIs, were involved in this study which had a duration from February 2020 to April 2020 and was approved by the ethical committee of the department of medical technology, university of Tripoli, Libya.
INCLUSION AND EXCLUSION CRITERIA
Patient's adults age ≥ 30 years with symptoms or suspected urinary tract infection (cystitis or pyelonephritis) or with a history of recurrent urinary tract infection were included in this study. Otherwise, Patients with asymptomatic bacteriuria and who have a catheter, urinary stent, or nephrostomy tube were excluded.
SAMPLE COLLECTION AND SAMPLING TECHNIQUE
Early morning mid-stream urine samples were collected using the sterile plastic disposable container and immediately transferred to the laboratory for investigation. Proper sampling instructions were given to each patient. [13] The collected urine specimens were subjected to general urine examinations using direct microscopy for white blood cells (WBCs) counting, then were inoculated on four types of media: Blood agar, Mannitol salt agar, MacConkey agar, and Cystine lactose electrolyte deficient agar CLED plates [14]. All inoculated plates were then incubated at 37° C for 24-48 hours for visible growth. Urine samples showing a colony count of more than 105cfu/mL were considered as positive pathogenic. The purified bacterial colonies were identified by gram staining, conventional biochemical procedures followed by a rapid biochemical test kit (API 20E). The Antimicrobial susceptibility test was performed using the modified Kirby-Bauer disc diffusion method according to the Clinical and Laboratory Standards Institute [15]. The following antibiotics were used: Sulfamethoxazole (SXT; 25 μg), Nitrofurantoin (F; 300 μg), Amoxicillin (AML;25 μg), Coamoxclav (AMC; 30 μg), Tetracycline (TE;30 μg), Ciprofloxacin, (CIP; 5 μg), Doxacycline (DO; 30 μg), Metronidazole (MTZ; 2.5 μg), Ceftriaxone (CRO;30 μg), Cefixime CFM; 5 μg). Isolates were classified as sensitive, intermediate, and resistant according to the standardized table supplied [16]. A sterile cotton swab was used to streak the surface of Mueller Hinton agar plates. Filter paper disks containing a designated concentration of the antimicrobial drugs were placed on the agar surface using an aseptic technique. Interpretation of antibiotic results was done by measuring the diameter of the zone of inhibition and compared to the table of antibiotic sensitivity. Zones of inhibition of more than 18mm were considered sensitive, 13-18 mm intermediate, and < 13 mm resistant [14]. Multi-drug resistant (MDR) strains were defined as those which showed resistance to three or more of the tested antibiotics.
RESULTS
Out of 200 urine specimens were included, of which 110 were female and 90 were male aged between 30-70 years old. However, only 110 (55.00%) of the total samples were positive for bacterial growth, 22 (20.00%) were male and 88 (80.00%) were female while the other 90 (45.00%) samples had shown no bacterial growth.
Gram-negative bacteria had a higher frequency of occurrence than gram-positive constituting 67 (76.14%) in females while 21(23.86%) in males from total culture.
The most frequent Gram-negative isolates organisms were Escherichia coli (49, 55.68%), the second most prevalent isolate was Klebsiella pneumoniae 18, 20.46%) followed by Pseudomona aeruginosa (9, 10.23%), Proteus mirabilis (8, 9.09%), Enterobacter aerogenes (2, 2.27%), and Citrobacter freundii (2, 2.27%). Meanwhile, the commonest isolate gram-positive bacteria were Staphylococcus aureus (20, 90.91%) followed by S. saprophyticus (2, 9.09%) (Table 1).
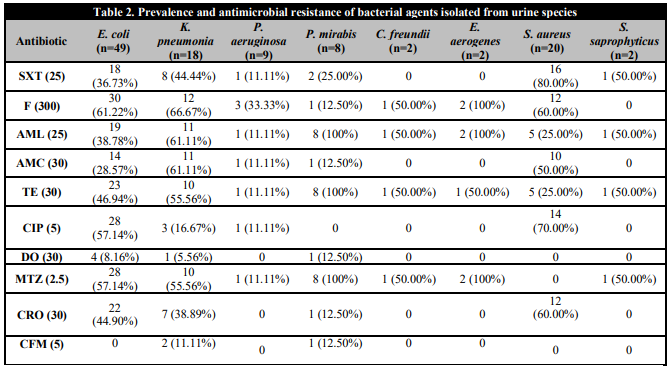
It was observed that Escherichia coli strains displayed relatively high antimicrobial resistance rates to Nitrofurantoin (61.22%), Ciprofloxacin (57.14%), Metronidazole (57.14%), Tetracycline (46.94%), and Ceftriaxone (44.90%) whereas it was least resistant to Doxacycline (8.16%). Klebsiella pneumonia were resistant to Nitrofurantoin (66.67%) followed by Amoxicillin (61.11%), Coamoxclav (61.11%), Tetracycline (55.56%), and Metronidazole (55.56%).
Among the Gram-positive isolates, S. aureus exhibited utmost resistance (80.00%) against Sulfamethoxazole, Ciprofloxacin (70.00%), Nitrofurantoin (60.00%), Ceftriaxone (60.00%), and had the lowest (25.00%) resistance to Amoxicillin and Tetracycline. S. saprophyticus presented moderately resistance to rate (50.00%) against Sulfamethoxazole, Amoxicillin, Metronidazole (Table 2). All other isolates were resistant to at least three antibiotics.
DISCUSSION
The misuse of antimicrobials to treat infections has a negative consequence on the organization of public health, as well as on the economic impact and the increase of drug resistance among causative bacteria. As a result, it is critical to continuously assess the state of antimicrobial resistance in a community, which was the study's first goal, especially in the case of UTIs.
The isolates obtained from UTIs in the current study demonstrated high resistance to different types of antibiotics. Most strains showed resistance to at least three antibiotics. Antibiotic resistance is a common phenomenon in developing countries such as Libya where drugs are available freely without prescription.
In the present investigation, E. coli is the important uropathogen causative, displayed a similar resistance pattern as for Klebsiella, and was showed resistance to Nitrofurantoin, Tetracycline, Amoxicillin, and Sulfamethoxazole/trimethoprim. This finding is inconsistent with the result obtained from previously reported in Libya which declared lower resistance rates for mentioned antibiotics [17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29]. We also found that most uropathogens exhibited high sensitivity to Cefixime followed by Doxacycline which was in disagreement with other studies' outcome in Libya [17, 18, 19]. The drug susceptibility profile of Gram-negative and Gram-positive bacteria tested in the present study was variable. For instance, increased bacterial resistance to ciprofloxacin has been shown. This study is opposite with the results reported by Elsayah et al [20] who revealed that Ciprofloxacin was the most effective antimicrobial agent. Therefore, we observed a higher frequency of ciprofloxacin-resistant in E. coli (57.14%) when compared to K. pneumoniae (16.67%). Meanwhile, S. aureus isolated strain also indicated higher resistance to ciprofloxacin (70.00%). The Specific ciprofloxacin resistance is also associated with chromosomal mutations altering DNA gyrasa and topoisomerase IV, overexpress efflux pumps, changing the number of porin sort, and transferring resistance to the plasmids' genes [21, 22]. Besides, most Gram-negative bacteria isolates in this study were resistant to three or more drugs [multi-drug resistance], as has been documented in other studies around the country [18, 19]. This demonstrates that multi-drug resistance is becoming a major issue in the treatment of uropathogens in Libya. This raises the need for national antimicrobial surveillance and in-vitro susceptibility testing program and strict adherence to antibiotic policy to prevent the spread of drug-resistant microbes in the region.
The gram-negative bacteria were the most common isolates in the current study, obtained in the present study was different in rates with other reports from different areas [18, 19, 23, 24]. This study detected the dominance of Escherichia coli (55.68%) and Klebsiella pneumonia (20.46%) (Table 1), which was almost identical compared with other research in Libya and other countries. In Northwest Libya, Abujnah et al have found a predominance of Escherichia coli (56%) and Klebsiella pneumonia (19%) [17]. In another study in Messalata, Libya, Mahammed et al have reported the predominance of Escherichia coli (56%) and Klebsiella pneumonia (17%) [18]. In Southern Tunisia, the authors have found Escherichia coli (68%) and Klebsiella pneumonia (13%) as predominance uropathogens among patients of UTIs [25]. A study in Iran has reported uropathogens with a predominance of Escherichia coli (38%) and Staphylococcus spp (35%). [26]. Staphylococcus aureus and S. saprophyticus were the most dominant Gram-positive uropathogen isolated in our study. Unlike in other studies which isolated coagulase-negative Staphylococcus and Enterococcus as the most dominant Gram-positive uropathogen [27, 28].
Our study also showed a high prevalence of UTI in females than males 88 (80.00%) and 22 (20.00%) respectively which correlates with findings from other studies which revealed that the frequency of UTI is greater in females as compared to males [13, 29]. This result is also in agreement with what was previously reported by Mahmoud and colleagues in 2016. Many other researchers have also reported similar findings [2, 8, 30]. The reason behind this high prevalence of UTI in females is due to the proximity of the urethral meatus to the anus, shorter urethra, sexual intercourse, incontinence, and bad toilet [29]. Nowadays, numerous organizations and programs are working to fight against antibiotic resistance [31] but the first step to obtaining proper management and good control policy for decreasing the development of antibiotic resistance among microorganisms, particularly the pathogens is the evaluation and practical assessment of the antibiotic resistance patterns among definite populations of the patients of a country.
CONCLUSIONS
In conclusion, we evidenced that E. coli was the most prevalent isolate followed by Klebsiella pneumoniae. These two organisms were highly resistant to the commonly used antibiotics. In addition, these organisms exhibit resistance too many first-line drugs used for UTI infection. To prevent resistance to antibiotics, appropriate therapy as per bacterial sensitivity pattern needs to be initiated.