INTRODUCTION
Lifestyle and dietary habits are major contributors to the growing incidence of non-communicable diseases. As a consequence of a dramatic increase in risk factors such as obesity, type-2 diabetes and altered food supply and preferences, non-alcoholic fatty liver disease (NAFLD) is one of the most important causes of liver disease worldwide, affecting an estimated ≥ 24 % of most populations (1). Genetic predisposition must be also placed in the context of environmental factors. Nowadays, accumulating evidence describes a ‘lean' manifestation of NAFLD, which was initially described in Asian populations and affects at least 5 % of the population in western societies (2). This ‘lean NAFLD phenotype' represents, among others, a ‘metabolically obese, normal weight phenotype' (1,2).
A significant body of data suggests that dietary composition and its interaction with the gut microbiota - and eventually the immune system - may predispose individuals to NAFLD, rather than total calorie intake, with a continuous positive balance promoting obesity, fatty liver, and finally NASH and/or metabolic syndrome. Over the past ten years, different mouse models - germ-free, antibiotic-treated and with or without colonization by specific microbial strains (3 4-5) - have been used to prove metabolic and immune effects derived from microbes for hosts. Here, imbalances in the gut microbiota have been linked to major risk factors for NAFLD (6,7). However, direct evidence for an association between functional alterations in the gut microbiota composition, as well as the microbiota-related molecular signature, and NAFLD and NASH remains largely inferential (3,5). In this scenario, surrounding environmental conditions, including diet (within 24 h), have proven to significantly impact microbial genome (microbiome) influencing host metabolic stress and dietary energy expenditure (8). Otherwise, long-term dietary habits were more clearly associated with imbalances in microbial levels categorized into enterotypes. The metabolic activity of gut microbiota has led to consider it as a ‘supraorgan' likely comparable to the liver. Notably, gut microbiome exhibits high variability between individuals (i.e., genders) and may constitute a source of variation for the effects observed, but evidence at the population level is incompletely characterized.
The liver is a central organ for the turnover and transformation of the metabolites supplied from the intestine. In addition, the intestine constitutes an important source of immune signals that stem from the intestinal mucosa and immune system. This so-called ‘gut-liver' axis is sensitive to the gut microbiota, which modulates energy supply and homeostasis, ectopic fat deposition, intestinal and fatty tissue inflammation, and immune maturation (9 10-11). The regulation of the energy balance operates both in the central nervous system and at peripheral sites, including adipose tissue and the liver, playing a critical role in the development of NAFL/NASH and liver fibrosis (12,13). In this sense, contradictory findings have been reported suggesting an important role for the diversity of gut microbiota as well as its composition, worsening or improving, both qualitatively and quantitatively, liver lipid accumulation (14,15). Besides, only a few studies have investigated the role (bidirectional) of the gut microbiota in host lipid metabolism (16) to establish steady-state commensalism and tissue metabolic homeostasis (17).
Preclinical data also support the relevance of pre- and post-natal nutrition to the developmental programming of NAFLD (18). The importance of perinatal outcomes in microbiota modulation at critical periods for development have been highlighted for their potential role in the prevention (19) and treatment of human disease. This is increasingly thought to occur through epigenetic changes as integral underlying mechanisms. Here, it can be established that an indirect gut microbiota-mediated epigenetic regulation may modify host immunometabolic responses and, ultimately, gene expression (20,21). These are novel perspectives connecting the intrauterine milieu to risk factors for NAFLD, i.e., diabetes, obesity, and hyperlipidemia, as well as for metabolic syndrome as a whole. However, how immunonutritional food components influence these aspects in NAFLD remains poorly understood.
In relation to the above, several recent reviews (22,23) have focused on reporting experimental and clinical evidence of intestinal dysbiosis in NAFLD. From these studies, conclusions were drawn on the therapeutic potential of manipulating the gut microbiota to decrease the incidence and prevalence of fatty liver disease. However, relatively few protocols have been registered in relation to gut microbiota manipulation in NAFLD patients (24). There is still an open debate about how and what microbes or their genomes may modulate NAFLD severity and (consequently) energy expenditure or nutrient fate to prevent metabolic and immune complications. Currently obtained results are promising, but as yet they do not permit definitive clinical conclusions in relation to their clear efficacy in microbiome-targeted therapies (24,25).
This review provides a comprehensive overview about the role of the gut microbiota as an important contributor to immunometabolic factors influencing liver disease. To this end a comprehensive update of the dynamic interplay between diet, nutrients, and microbes, as well as the microbial phylogenetic changes that occur in obese versus lean individuals is reported. Particularly, this manuscript is focused on the immune function and includes the current knowledge on the functional outcome obtained from modifications in microbial ecosystems.
EXPOSOME AND LIVER FUNCTION
The mammalian gut harbours highly complex microbial communities, which develop multifaceted interactions in the health and nutrition of the host and are influenced by the exposome (i.e., environmental exposures to various exogenous - diet - and endogenous factors), and have influential effects on both metabolism and immunity. Microbial products derive from either saccharolytic or proteolytic fermentation and may affect the gut-liver axis via multiple mechanisms, hence contributing to the pathogenesis of NAFLD.
Different microbial-mediated processes have received major attention, including: a) efficiency of energy harvest, i.e., utilization of nutrients (fermentation of complex dietary carbohydrates) favouring short-chain fatty acid (SCFA) production (5); b) modulation of low-grade liver (and adipose tissue) inflammation, e.g., via activation of free fatty acid receptor 2 (FFA2, a G-protein-coupled receptor also known as GPR43) (26); c) impact on the innate immune response, such as endotoxin production from Gram-negative bacteria (27), and the adaptive immune response, by binding to leukocyte receptors (butyrate binding to GPR41 and GPR43) suppressing lymphocyte proliferation, to inhibit cytokine production by Th1-lymphocytes and to induce T-lymphocyte apoptosis (28).
SCFAs can regulate lipogenesis through their interaction with the endocannabinoid system or stimulation of orexigenic hormone release (e.g., leptin) in adipose tissue (29). Also, they can reduce or increase insulin resistance (30) via a GPR40-mediated and calcium-dependent mechanism regulating the expression of PPARδ. Otherwise, the impact or modulatory role of bacterial metabolites originating from a proteolytic metabolism on host physiology has received much less attention.
Microbes gradually shift to their proteolytic metabolism as carbohydrates are depleted in the intestinal lumen. Bacterial protein degradation products are considered deleterious for the host's physiology due to the production of phenolic compounds (i.e., indole) derived from aromatic amino acid degradation. Numerous Gram-negative, as well as Gram-positive, bacterial species able to produce indole have been identified (including Escherichia coli, Clostridiumspp., and Bacteroidesspp), including Lactobacilliand Bifidobacteriathat are considered beneficial for the host's health. Accumulating evidence suggests that microbial tryptophan catabolites resulting from proteolysis have an impact on mammalian gut immune homeostasis through binding to the aryl hydrocarbon receptor (AHR), stimulate hormone (i.e., GLP-1) secretion, and modulate gastrointestinal motility (31). Furthermore, adequate tryptophan metabolism can prevent redox imbalances due to decreased production of NAD equivalents, which are used to compensate the fermentation of carbohydrates.
Nutritional deficiency of choline has also been associated with changes in the microbiome of the human gut and the development of fatty liver, as previously found in rodent models of NASH (32). Accordingly, changes have been reported in various physiological processes such as lipid metabolism, enterohepatic circulation of bile acids, and cholesterol metabolism (33). These observations were accompanied by increased numbers of the Gram-negative bacteria Gammaproteobacteriaand Erysipelotrichi, whose presence has been associated with increases in fat content in the liver (34). The Erysipelotrichaceaefamily is relatively distant, from a phylogenetic point of view, from the Clostridialesorder, for which oxidative enzymes contributing to the production of oxysterols have been identified. This bacterial metabolic potential opens new avenues to the microbial contribution to synergistic lipotoxic effects leading to mitochondrial function impairment (35).
Collectively, existing data demonstrate that the gut microbiome can affect the liver, the adipose tissue, and metabolic diseases both in a beneficial and in detrimental way. However, owing to the complexity and multifaceted nature of the interactions between gut microbiota and its microbiome with the host's nutritional and health status, no human in vivodata on a direct link between microbial-derived metabolites, liver function, and NAFLD progression are available.
ENERGY HARVEST AND FATE OF NUTRIENTS
The commensal microbiota regulates nutrient acquisition and utilization in symbiosis with the host, providing additional enzymes (36,37). Here, it is implied a regulated expression of various genes that are involved in the utilization of dietary carbohydrates and lipids. Metagenomic studies identified gut microbial enzymes specialized in the utilization of otherwise non-digestible nutritional carbohydrates and host-derived glycoconjugates (e.g., mucins, proteoglycans). These genes participate in the deconjugation and dehydroxylation of bile acids, in cholesterol reduction (the latter being mainly converted to coprostanol by intestinal fermentative bacteria, either directly by the enzyme cholesterol reductase or via an alternative indirect pathway), in the biosynthesis of vitamins and isoprenoids, and in the metabolism of amino acids and xenobiotics (36,38).
Thus, novel human microbiome-targeted interventions have been proposed, while an effective approach has yet to be successfully developed (24). Human trials support potential positive roles for prebiotics (i.e., non-digestible food ingredients, mostly oligosaccharides that stimulate growth and activity in several specific, beneficial intestinal bacteria) enhancing NAFLD/NASH progression likely through their modulatory effects on glucose homeostasis and lipid metabolism (5). Also, the few existing studies revealed changes in the microbiome after very short-term dietary interventions (10 days) with high-fat/low fibre versus low-fat/high fibre (39). However, direct evidence of the influence of defined prebiotic structures on specific microbial genes and their physiological functions is lacking. Nowadays, prebiotic contribution to establishing a ‘score' of disease and validating its use in clinical practice remains elusive.
The potential contribution of microbial fermentation has also been a matter of controversy since it improves energy harvest from the diet, which may contribute as much as 10 % to daily calorie intake (40). Studies in rodents clearly showed the causative role of the gut microbiota in adiposity and insulin resistance, linking changes in host phenotypes to variations in gut microbial composition (14, 41, 42, 43). Thus, germ-free animals did not develop diet-induced obesity and related comorbidities, like visceral adipose tissue inflammation with macrophage infiltration and insulin resistance (42). Otherwise, these mice gained excess fat mass when conventionalized with microbiota from lean non-germ-free animals kept on a normal diet (14). The persistent lean phenotype of the germ-free animals displayed increased skeletal muscle and liver levels of phosphorylated AMP-activated protein kinase (AMPK), accompanied by its downstream targets involved in fatty-acid oxidation (acetylCoA carboxylase, carnitine-palmitoyl transferase) (14). Fat mass gain was greater when microbiota from ob/obmice (obese due to defective leptin signalling) was transferred to germ-free animals (41). The ob/obmicrobiome showed a higher proportion of environmental gene tags encoding many enzymes involved in breaking down otherwise indigestible dietary polysaccharides, including pathways for starch/sucrose, galactose and butanoate metabolism, indicating an improved energy harvest (41). In these scenarios, a comparison of microbial families associated to diabetes-related clinical traits revealed a positive correlation with Lactobacillaceae, while Bacteroidaceaewas negatively associated with weight gain (43). Dietary supplementation (12 weeks) with prebiotic galacto-oligosaccharides (GOS) was proven effective to increase fecal numbers of Bifidobacteriumspp. without improving insulin sensitivity in overweight or obese prediabetic men and women (44). Metagenomic studies in patients with T2D revealed that changes in the microbiome rather than in microbial taxa have a greater influence on the risk factors for T2D (45).
MICROBIOTA IN METABOLIC AND ENDOCRINE PROCESSES
Promising reports on the effectiveness of microbiome-based targeted interventions demonstrated changes in hepatic gene expression when animals were fed prebiotics (46,47). Gene expression implied that different genes are involved in diverse biochemical processes: lipogenesis, cholesterol production and mobilisation, or induction of PPARα-mediated fatty-acid oxidation. Notably, the composition of the gut microbiota, including the activity and growth of lactobacilli and bifidobacteria, which promote the production of SCFA (e.g., acetate, butyrate and propionate), appeared as a critical factor driving the magnitude and final biological outcomes. Apart from trophic effects on intestinal epithelia (48,49), reported findings remark the high dependence of biological effects in the host's physiology on the SCFA profile produced. While the production of luminal acetate appears to be a precursor for hepatic lipid synthesis (16), propionate oppositely impacts lipid metabolism inhibiting the effects of acetate. Generally, butyrate is regarded as a healthy metabolite as it is utilized by enterocytes, positively influencing cell growth and differentiation, and promoting anti-inflammatory effects. Collectively, it was found that increasing proportions of SCFA producers significantly alleviate hepatic steatosis, decreasing the content of triglycerides and free fatty acids in rats fed a high-fat diet (50). The latter could open new interpretations to the role of lactobacilli, major contributors to the efficiency of energy harvest and nutrient utilization, and their potential contribution to the onset and progression of NAFLD. SCFAs also affect endocrine functions due to their interaction with G-protein-coupled FA receptors (i.e., GRP43 and GRP41) (51), the stimulation of which increases the production of peptide YY and leptin (orexigenic and anorexigenic peptides, respectively) (52,53).
Much of the beneficial effects of prebiotics on plasma lipid and hepatic triglyceride concentrations are attributed to regulation of the expression of the different transcription factors involved in lipid metabolism (i.e., acetyl-CoA carboxylase, fatty-acid synthase, malic enzyme, and ATP citrate lyase). These effects occur via regulation of glucose metabolism (54) and insulin resistance (55). This could point to enhanced enteroendocrine function in L-cells (mainly located in the ileum and colon) (56) as mediators of the trophic effects associated with reduced mucosal inflammation (48,57) and induction of the antidiabetic (PPAR-mediated regulation of Akt pathway influencing GLUT4 expression in liver) and anorexigenic incretin glucagon-like peptide 1 (GLP-1).
Prebiotics have also been proven to influence physiological endocrine systems up-regulating the production of GLP-2 and expression of the endocannabinoid system (eCB) (58). Thus alleviating gut permeability, plasma lipopolysaccharides (endotoxaemia) from Gram-negative bacteria (LPS) and cytokines decreased the hepatic expression of inflammatory cytokines (e.g., TNF-α, MCP-I) and oxidative stress markers in obese (leptin-deficient) mice. It has been recently shown that a dysregulated expression of eCB 1 and 2 in the colon and liver is associated to metabolic imbalances derived from sirtuin 3 (SIRT3) deficiency (59). These observations were followed by increased endotoxaemia in correlation with gut microbiota dysbiosis. Yet the underlying mechanism linking SIRT3 with gut microbiota remains obscure, and SIRT3 deficiency may be attenuated by sodium butyrate. Prebiotic-treated obese (ob/ob) mice exhibited a normalized eCB system tone (expression of eCB-1r, plasma eCB levels or adipose tissue eCB content) in both the gut and adipose tissue, as well as endotoxaemia (29). Figure 2summarizes the interrelationship between prebiotics, gut permeability and eCB-induced adipogenesis and NAFL.

Figure 2. Interrelationship between prebiotics, gut permeability, and influence of the endocannabinoid (eCB) system in adipogenesis and NAFL development.
Relatively rapid shifts in gut microbiota composition and their related functional outcomes in response to high-sugar and high-fat diets have been reported (8,60,61). These responses modify both gene expression and the resultant metabolite generation and flow in the host. At the same time, alterations in the composition of the gut microbiota influence strain-specific susceptibility to diet-induced metabolic disease (43). This bidirectional interaction determines the microbial capacity for processing dietary sugars and for generating hydrophobic bile acids and, thereby susceptibility to metabolic disease. The impact of metabolic disease on bacterial functional outcomes is reflected, whether worsening or improving the disease, in the role of GPR43 as sensor for excessive dietary energy (15). Apparently, this receptor loses its inhibitory influence on insulin signaling, reducing fat accumulation in adipocytes. An example is vancomycin-treated mice, which display microbial changes relatively similar to obese patients, with a significantly reduced diversity of the gut microbiota and a markedly increased abundance of Lactobacillusspp (62). However, these bacterial changes were accompanied by significant increases in body weight and colonic GPR43 expression. Microbial alterations in obese people (60,63,64) favor higher glycoside hydrolase activities and a higher activity of host transport proteins importing the breakdown products of indigestible carbohydrates.
MICROBIOME AND LIVER FUNCTION
Grain-based food, sugars and dairy products are the nutritional staples of western diet, which produce high glycemic and fat loads that have been recognized as aggravating factors of fatty liver disease.
Western diets, with the concomitant specific changes in gut microbiota to pro-inflammatory microbes (Gammaproteobacteriagenera including Klebsiellaspp., Enterobacterspp., and Escherichiaspp.), also reduce the availability and utilization of several amino acids originating from both alimentary and endogenous proteins. These metabolic conditions favour the development of fatty liver inflammation in patients on controlled diets with lowered levels of choline (32,65). Dietary choline deficiency is an example of one (of several) nutritional factors in modern diets that may favor the development of NAFL and NASH. Most studies documented that choline is an essential nutrient with a key role on fatty liver progression by decreasing methionine levels, favoring abnormal phospholipid synthesis and lipoprotein secretion, mitochondrial dysfunction, and endoplasmic reticulum stress (32,33). In addition, choline deficiency contributes to dysregulate methionine levels and tissue polyamines, a significant part of which have their origin in the intestinal microbiota, impairing glucose, lipid, and energy homeostasis.
Studies correlating the content of polyunsaturated fatty acids (PUFA) with the development and extent of NAFL have produced divergent results (66 67-68). Dietary deficiency of ω-3 PUFA caused imbalances on microbial composition in mice, with decreased cecal numbers of Lactobacillusspp. and Bacteroidetes-Prevotellaspp. (67). However, ω-3-depleted mice displayed a slightly increased proportion of Bifidobacteriumspp., which was significantly associated with increased insulin resistance and steatosis. Here, dietary supplementation with fructo-oligosaccharides (FOS), for only 24 days, increased the proportion of lactobacilli, contributing to restore an efficient fat storage without lessening liver steatosis (67). Thus, these studies demonstrate different effects of prebiotic fibres in fatty liver disease depending on dietary microbiome adaptations. Also, the results may be interpreted to suggest a direct role for microbial fermentation in steatosis, insulin resistance, and down-regulation of hepatic inflammatory regulators (i.e., peroxisome proliferator-activated receptor (PPAR)-α, and receptor gamma coactivator α). Collectively, it can be delineated a significant bulk of evidence supporting the bidirectional impact of either nutrient availability or gut microbial imbalances on each other to trigger profound effects on mammalian lipid metabolism. Table Isummarizes the effects observed in serum and liver lipids, as well as in the biomarkers associated to NAFL, in different animal models fed a high-fat diet.
Table I. Examples of the effects of prebiotic fibres on serum and liver lipids and biomarkers associated to NAFL in animal models fed a high-fat diet

Indirectly, gut microbiota can modify the lipid profile in the enterohepatic circulation affecting overall hepatic lipid metabolism. Solely microbial transformation of cholesterol and bile acids constitute the origin of coprostanol and secondary bile acids - deoxycholic acid (3α,12α-dihydroxy-5β-cholan-24-oic acid) and lithocholic acid (3α-hydroxy-5β-cholan-24-oic acid). Under physiological conditions these bile acids are absorbed but human hepatic metabolic capacities cannot remove them from the bile acid pool causing hepatic toxicity (69). 7α-dehydroxylation is the most quantitatively important bacterial bile salt biotransformation in the human colon, and 16S rDNA phylogenetic analyses have led to classify most competent intestinal bacteria into the genus Clostridium(70). The order Clostridiales, including ClostridiumXIVa and IV subgroups, within the superfamily Lachnospiraceaeis peripherally related to the family Erysipelotrichaceaewithin phylum Firmicutes(71) mostly associated to obese individuals (72). Unfortunately, there are scarce studies addressing to what extent these differences in hydroxysteroid dehydrogenases could be responsible, at least in part, for the significant detrimental effects in NAFLD as 7β-dehydroxylation activity has also been described in Bacteroides distasonis(73).
Despite the direct effect of bile acids helping in the intestinal absorption of lipids and fat-soluble vitamins, they also play a role in regulating lipid-glucose homeostasis, inflammation, and thereby innate immune cells polarization via specific nuclear receptors (FXR) and G-protein-coupled receptors (TGR5). FXR is highly expressed in the liver, intestine, kidney and adrenal gland, and may play a role in reducing hepatic triglyceride levels via different gene pathways, including FGF21 (74), ChREBP (75), PPARα (76), and AKR1B7 (77). TGR5 is highly expressed in the gallbladder, intestine (primarily in the ileum and colon), and liver, where it has been shown to antagonize the activity of NFkB in macrophages and hepatic Kupffer cells (78). Moreover, in intestinal L-cells TGR5 induces GLP1 secretion, and in the liver TGR5 attenuated triglyceride accumulation through a yet-to-be-determined mechanism (79).
MICROBIAL IMBALANCES ASSOCIATED WITH LIVER DISEASE
A direct link between NAFL/NASH status and imbalances in the composition of the gut microbiota is now being established (3,27). The most relevant alteration is an increased ratio of Firmicutesto Bacteroidetes, associated to the overgrowth of Gram-negative bacteria and decreased proportions of Bifidobacteriumspp. in the cecal contents of obese humans (63,80). However, contrasting results have also been obtained from clinical studies where the proportion of Bacteroideswas found significantly increased in subjects with liver fibrosis (81). In a similar way to the procedures followed to uncover the role of gut microbiota in obese and T2D patients, ‘infection' models served to establish the microbial contribution to the induction of adaptive and innate immune memory (82). This bacterial contribution seems to occur at the epigenetic, metabolic, or functional level. However, despite its potentially important role, to the best of our knowledge no population-based evidence is available regarding the ability of the mucosal immune system to co-operate with systemic immunity in response to certain commensals in the setting of NAFL.
Scientific evidence available in the literature tried to establish a relationship between bacterial translocation through the intestinal barrier and NAFL aggravated symptoms and disease progression (83,84). The overgrowth of small intestinal bacteria (SIBO) is directly associated with an increase in intestinal permeability affecting nutrient absorption and utilization. Notwithstanding this, the impact and magnitude of intestinal functional impairment seem to be associated to the NAFLD phenotype.
For example, a study population (n = 35 patients) with biopsy-proven NAFLD determined the presence of a "leaky" intestine with evidence of increased SIBO (83). Furthermore, a cross-sectional study performed in outpatients with metabolic risk factors for NAFLD development (type-2 diabetes, obesity, dyslipidemia) did not find any significant association between SIBO and NAFLD or hepatic steatosis (85). However, it was found that central obesity was significantly associated with SIBO.
Metagenome-wide association studies revealed that gut microbiota in T2D patients was characterized by decreased numbers of some universal butyrate-producing bacteria (Eubacterium rectale, Faecalibacterium prausnitzii, Roseburia intestinalisand R. inulinivorans). Besides this, an increase in various opportunistic pathogens (Bacteroides caccae, Clostrium hathewayi, Clostridium ramosum, Clostridium symbiosum, Eggerthella lenta, and Escherichia coli) was quantified, as well as an enrichment of other microbial functions conferring sulphate reduction and oxidative stress resistance. Obesity was associated with reduced bacterial diversity, with lower numbers of Bacteroidetesand increased numbers of Actinobacteria(30,63,86). Some other components of the obesity-associated gut microbiome are distributed among Proteobacetria, Fusobacteriaand Verrucomicrobia(72). Otherwise, contrasting data are reported in relation to the proportion of Methanobrevibacter smithiiin obese subjects (63,72). Unexpectedly, M. smithiiand Escherichia coliwere found to negatively correlate with body mass index in overweight and obese people (87). Recent extensive studies have revealed that molecular hydrogen (H2) has a great impact on obesity development with a decreased expression of H2S-synthesizing enzymes and reduced H2S levels in adipose tissues in mice. Also, significant increases in some other bacterial groups such as Staphylococcus aureus(88,89) and Faecalibacterium prausnitzii(90) have been associated with NAFL comorbidities. However, a decreased abundance of both the Faecalibacteriumand Ruminococcusgroup was recently demonstrated in ‘lean' NASH patients (91).
The role of Lactobacillusspp. in obesity has been a subject of controversy, with part of the risk for causing public health problems in humans being attributed to the promotion of consumption of products containing these bacteria. Analyses on the proportions of Lactobacillusspp. evidenced contrasting changes in this microbial population. While significant increases (63,92), particularly in the species L. reuteriand L. ingluviei(63,93), have been found in overweight people, significant reductions in the fecal concentrations of the Lactobacillus gasserisubgroup were observed in obese infants at baseline (80). Nowadays few interesting data are available regarding the role of dietary supplementation with probiotics in the management of obesity, as most probiotics for human consumption belong to the Lactobacillusgenus. In this sense, a recent study in obese children who received daily supplementation with L. casei(Shirota strain) microbial effectiveness contributing to weight loss while improving lipid metabolism was demonstrated. Notably, these effects could be associated to a significant increase in the fecal numbers of Bifidobacteriumspp. and acetic acid concentration (80). These results suggest that microbial metabolic associations are important to determine the functional outcomes of gut microbiota composition in the development of metabolic diseases. Table IIsummarizes the most significant bacterial groups associated with overweight in obese animals and humans. The specific role of gut microbiota components and their bioactive factors remains unclear in NAFLD. In addition, the direct role of gut microbiome in the development of obesity and its associated phenotypes has been demonstrated to be transmissible by fecal transplantation into germ-free mice (43,94). Here, bacterial colonization in obesity and associated disorders appears to be dependent on host metabolic responses to diet.
Table II. Summary of the most significant bacterial groups associated with overweight and obese animals and humans

In addition to the gut microbiota metabolic participation in the modulation of lipid metabolism, it has also been associated with an indirect regulatory influence in the central nervous system (CNS), the so-called ‘gut-brain' axis. Thereby, microbial-mediated CNS control could influence food intake and expenditure, the regulation of hepatic lipogenesis, and gut integrity, as well as hepatic low-grade inflammation. Preclinical studies have shown that microbial colonization of germ-free mice can reduce the levels of circulating fasting-induced adipose factor (Fiaf), and the levels of phosphorylated AMP-activated protein kinase (both in the skeletal muscle and liver), thus contributing to fat storage (7,95) (Fig. 1). Taken together, these functional outcomes of gut microbiota beyond energy harvest from diet have led to the concept that gut microbiota serves as an important regulator of energy homeostasis and ectopic fat deposition. Nevertheless, the functional characterization of transplanted microbial communities suggest that variation in metabolic outcomes is under the control of both host genetics and gut microbiota (43).
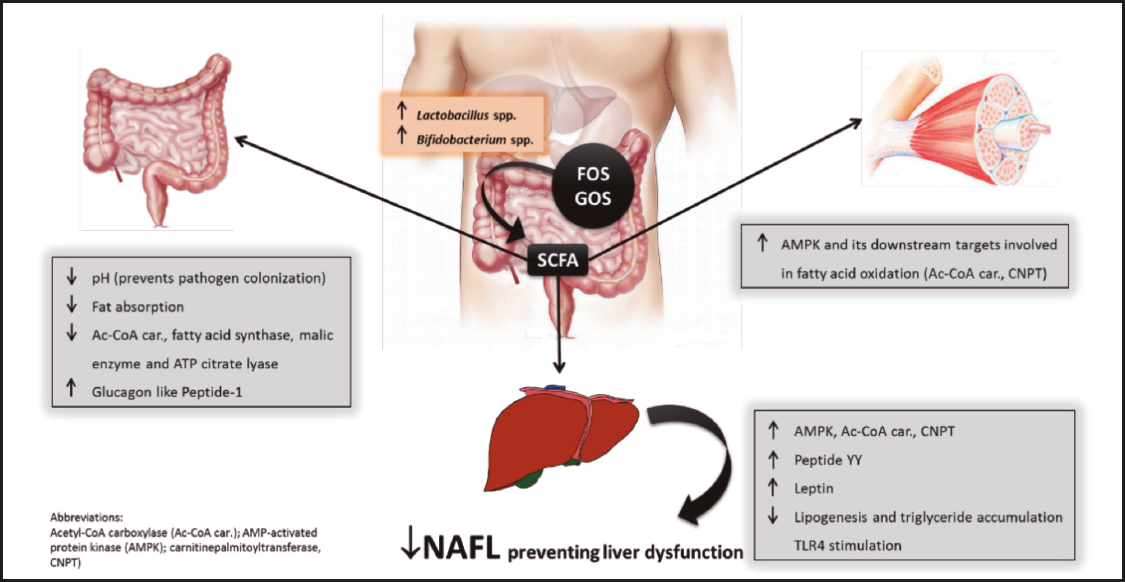
Figure 1. Beneficial effects of short-chain fatty acids (SCFA) derived from bacterial fermentation of dietary fructo-oligosaccharides and galacto-oligosaccharides on non-alcoholic fatty liver (NAFL) development.
Data from scarce nutritional interventions in humans also strongly suggest the microbiota-mediated positive effects in patients with nonalcoholic steatohepatitis (NASH) (96). A study in 66 patients who received a symbiotic mixture of Bifidobacterium longumwith FOS for 24 weeks concluded that, compared to lifestyle modification alone, dietary supplementation with the symbiotic formulation significantly reduced NASH activity index, serum concentrations of LDL cholesterol, inflammatory markers, and insulin resistance. Otherwise, knockout mice for phosphatidylethanolamine N-methyltransferase (PEMT) develop severe NASH (97). However, these mice are protected from obesity and whole-body insulin resistance. Here, the study of the influence of feeding prebiotics to animals with metabolic risk factors for NAFLD development has been inferential. Feeding the prebiotic kojibiose (2-O-α-D-glucopyranosyl-α-D-glucopyranose) to hyperglycaemic animals treated with arachidic acid significantly reduced the plasma levels of lyso-phosphatidylcholine (98). These changes were accompanied by body weight loss and increased numbers of Bifidobacteriumspp. (by 12 %), Bacteroidesspp. (by 2 %), and Enterobacteriaceae(by 4 %) relative to control animals.
POTENTIAL DUAL ROLE OF PROBIOTICS
To date there is a controversial debate concerning the species- or strain-dependent role, whether positive or negative, of Lactobacillusspp. in body weight gain (92,99). Different investigations point out specific alterations in gut microbiota composition causing markedly opposite regulatory effects on lipogenesis associated with increased numbers of Lactobacillusspp., and thereby Firmicutesin fecal contents. This is of particular importance since most probiotics for human consumption belong to this genus. While some species (L. acidophilusand L. reuteri) have been associated with obesity-associated gut microbiota (63,64), others (L. plantarumand L. gasseri) are associated with weight loss in obese humans and animals (63). Feeding L. ingluievito mice preceded an increase in weight and increased FAS and TNF-α transcript levels in the liver (93). L. acidophilusor L. caseiwere likely protective against diabetes since animals fed with these probiotics did not show any evidence of metabolic syndrome such as insulin resistance (100). However, in some cases strain-specific beneficial effects have been reported, for example, in patients dietary supplemented with L. plantarum299v or a yoghurt supplemented with L. acidophilus145, B. longum913 and oligofructose (101,102). Most prominent effects significantly reduced serum concentrations of LDL-cholesterol (by 9.6 %) and fibrinogen (101) or increased HDL-cholesterol (by 25.8 %) (102). Feeding L. rhamnosusGG and L. sakeiNR28, supplemented within a normal chow, to mice resulted in a significant reduction of epididymal fat mass, as well as obesity-related biomarkers like AA, FAS, and steatoryl-CoA desaturase-1 in the liver (103). Taken together, all studies seem to highlight strain-dependent effects on lipid metabolism and liver/adipose tissue inflammation for Bifidobacteriumand Bacteroidesspp. that counteract NAFL/NASH.
Another important aspect is that most species of these genera possess β-glucosidase activity contributing to nutrient utilization. Otherwise, metagenomic studies have also demonstrated the expression of β-glucuronidase activity in some species of Firmicutes(104). β-glucuronidase activity might contribute to the production of toxins and/or mutagens resulting from increased intestinal reuptake and hepatic retention of usually deconjugated, potentially toxic bile acids. However, the role of bacterial β-glucosidases in the development of NAFL/NASH needs further study, since opposite bacterial patterns have been found as a function of the aetiology of fatty liver disease. For example, enteric dysbiosis with relative abundance of Bacteroidetesand Verrucomicrobiacompared to Firmicutes, a pattern that should correlate with amelioration of NAFL/NASH or obesity, has been associated with alcohol-induced steatosis and steatohepatitis (105).
INNATE IMMUNITY
NAFLD has been strongly associated with gut injury and increased intestinal permeability, and NASH frequently recurs in patients who underwent weight reduction surgery to treat obesity. These conditions favour translocation of bacterial antigens across the intestinal mucosal wall into the portal circulation in 25-70 % of patients with liver disease, enhancing hepatic inflammation and disease progression to NASH (9,11). Studies in animals and humans indicate that gut-derived bacterial endotoxin (i.e., lipopolysaccharide, LPS) and other cell wall components play important roles in the development of NAFL and its progression to NASH by stimulating, for instance, toll like receptor (TLR)-4. This receptor is prominently expressed on innate immune cells, increasing the release of proinflammatory cytokines and chemokines favouring insulin resistance (106).
Immune crosstalk between gut microbiota and its host takes place by interaction of the gastrointestinal epithelium and different cell types in the gut-associated lymphatic tissue that possess TLRs (107 108-109). These receptors recognize specific molecular patterns (proteins, nucleic acids, lipopeptides, etc.) found in both commensals and pathogens. Especially, recent reports highlighted the potential key role of defined Bidifobacteriumspp. (109) and Lactobacillusspp. (108) strains as promising modulators of inflammatory activity by regulating TLR-2/4 and NF-κB expression. Amelioration of LPS-induced NF-κB activation by potentially beneficial bacteria could, at least in part, explain their positive effects on the inflammatory signals originating in the intestine, preventing further amplification for those within the gut liver axis. The importance of innate immunity has been demonstrated in fructose-treated mice, as a model of steatosis characterized by high lipogenesis, where increased accumulation of 16S rDNA sequences in the liver was associated with significant induction of TLR 1-4 and 6-8 (34). However, mice genetically deficient in TLR5 also exhibit hyperphagia and develop hyperlipidemia, hypertension, insulin resistance, and increased adiposity (110). In contrast, TLR2-deficient mice under germ-free conditions are protected from diet-induced insulin resistance resulting in a phenotype reminiscent of metabolic syndrome (111). These animals were characterized by differences in gut microbiota, with a 3-fold increase in Firmicutesand a slight increase in Bacteroidetesas compared with controls, although both were present in similar proportions (47.9 %) (111). These authors suggested that the presence of gut microbiota could reverse the phenotype of an animal developing as a consequence of glucose intolerance and later obesity. Fructose disposition (solid or liquid formulations) has been shown to differentially affect gut microbiota composition and Bacteroidescolonization in mice (112). The decreased numbers of, among others, Bacteroidesspp. and Lactobacillusspp. were associated with up-regulated protein expression in the lipopolysaccharide-dependent proinflammatory TLR4/NLRP3 inflammasome pathway. Recovery of the intestinal Bacteroidetespopulation could contribute to better balance immunostimulatory thresholds because of the pivotal role played by the outer membrane vesicles (OMVs) of Gram-negative bacteria and their interaction with TLRs. Bacteroides uniformisCECT 7771 was reported to help in developing adequate immune responses, improving macrophage activity and liver steatosis (reduced fat vacuoles in liver tissue) in mice (113).
Studies conducted in rodent NASH models have demonstrated a direct implication of gut microbiota in enhancing NASH severity (114). When feeding a methionine-deficient diet for 4 weeks, animals develop defective NRLP3 and NRPL6 inflammasome (damage-associated tissue) sensing. This nutritional deficiency causes an aberrant colitogenic intestinal microbiota characterized by increased proportions of Bacteroidetes(Prevotellaceae) and bacterial phylum TM7, and reductions in the genus Lactobacillus in the Firmicutes phylum (115). This microbiota composition, although contrasting to that reported in obese and T2D patients, was identified as responsible for colonic inflammation (CCL5 secretion), favouring the translocation of TLR-4/9 stimulants towards the portal circulation (114). Similar inflammatory conditions were reported in fibrogenic animals administered CCl4 associated to bacterial translocation with increased proportions of aerobic bacteria and decreased numbers of Gram-positive anaerobics in the Clostridiumgroup (116).
Metabolic endotoxemia can result from multiple mechanisms, including increased production of endotoxins upon changes in gut microbiota composition, and triggers TLR4 receptor complex (i.e., TLR4/MD2/CD14) signaling in murine liver cells (mice fed a choline-deficient diet for 3 weeks) to fuel liver and adipose tissue inflammation (117). Thus, a functional relationship between metabolism and the innate immune system is increasingly recognized, suggesting that innate immune alterations can favour the development of metabolic syndrome (109,118). In this regard food ingestion, even in lean subjects, triggers a mild inflammatory response in the liver and adipose tissue, which usually does not lead to negative consequences. However, with the advent of immunometabolic imbalances such as obesity coupled with an unfavourable lifestyle and certain medications or genetic predisposition, inflammatory responses can get out of control, thus contributing to NASH development and severity (119). Accordingly, high calorie feeding induced a postprandial increase in surrogate serum markers of liver fibrosis in patients with NAFL and T2D (120). Here, inflammation is driven by classically activated (M1 polarized) macrophages, both in the liver and adipose tissue (121), where TLR4 receptor complex activity significantly contributes to macrophage polarization and functional differentiation (117,122). Thus, TLR4-/-mice fed a choline-deficient diet (for 22 weeks) displayed reduced hepatic macrophage infiltration with decreased macrophage chemoattractant protein (MCP)-1 and chemokine-chemokine receptor (CCR)-2 expression (123). Also, the hepatic proportion of F4/80+macrophages significantly increased in GPR43-/-mice fed a high-fat diet (15). Taken together, these studies highlight a direct immunometabolic control by microbial metabolites on the immune innate cell population of macrophages. In this regard, microbial butyrate has been proven to modulate macrophage metabolism, reducing mTOR kinase activity and increasing their antimicrobial phenotype (124).
HOST GENOME-MICROBIOME INTERACTION
Accumulated evidence regarding the immunometabolic implications of intestinal dysbiosis proved the (epi)genomic relationship between the exposome and host genetics to influence gut microbiome. The bidirectional interaction of gut microbiota and its microbiome with the host's physiological processes also induces changes in bacterial metabolism. This metabolic adaptation can lead to pattern changes in microbial metabolites, which may be important indirect regulators of the host epigenome, worsening or improving the development of liver disease. An example of this microbiome adaptation involves the reported changes in the gene composition of gut microbiota, which are similar to those seen in the gut microbiota of patients suffering from inflammatory bowel disease as compared to lean controls (86,125). Obesity and type-2 diabetes microbiomes present the hallmarks of adaptation to low microbial diversity environments (43,125). These microbiomes are enriched with genes encoding enzymes frequently involved in membrane transportation (125) and exhibit differences in glucose-stimulated insulin secretion (43). Furthermore, these microbiomes are depleted of cofactor, vitamin and nucleotide metabolism and transcription (125). Genomic studies have shown a strong influence of microbial metabolites (SCFAs) in the regulation of epigenetic programming in different critical tissues (liver, white adipose tissue, and proximal colon), determining the onset and severity of NAFLD (125). Collectively, these studies demonstrated that intestinal dysbiosis yields a core microbiome closely related at a functional level to systemic immunometabolic diseases and different physiologic states.
The pathogenesis and severity of NAFLD has been associated with several nonsynonymous single-nucleotide polymorphisms in the PNPLA3gene (rs738409, rs143392071) (127), increasing the risk for both steatosis and fibrosis (128); the MTTP (rs756998920) gene (129), and the adiponectin (rs1501299) gene (61). Also, a splicing mutation in TM6SF2 (rs58542926) affecting postprandial lipoprotein metabolism and glucose homeostasis in NAFLD has been identified (130). Research advances in this sense determined that the PNPLA3, TM6SF2, and MBOAT7 variants are associated with increased liver injury. The TM6SF2 variant seems to modulate predominantly hepatic fat accumulation, and the MBOAT7 polymorphism is commonly linked to fibrosis.
Data from animal models indicate that maternal overnutrition, even in the absence of pregnancy-related obesity, can promote metabolic dysregulation and predispose to T2D in the offspring (131). Metabolic programming of disease is being increasingly thought to occur via epigenetic changes as integral underlying mechanisms (132). Accordingly, epigenetic studies in diabetic and obese subjects revealed significant methylation differences in the promoter region of innate immune mediators such as TLR-2 and -4 (133), and FFAR3 (134) in comparison to lean controls, suggesting that changes in gut microbiota are involved in the epigenetic regulation of inflammatory reactions.
CONCLUSIONS
NAFLD has been revealed to be a severe liver dysfunction with an increasing prevalence in both obese and lean patients with metabolic syndrome where overnutrition and sedentary lifestyles are considered major contributors. There is a cumulative body of evidence for a correlation between alterations in microbial diversity and the functional output of gut microbiota. These changes appear as important modulators for the development of NAFLD or its progression toward NASH. Phylogenetic changes occur in the microbiota of obese versus lean individuals, and are characterized by an increased Firmicutesto Bacteroidetesratio. Here, the relationship between Lactobacillusspp. and Bifidobacteriumspp. appears to be critical for energy harvest and endocrine regulation. Thus, it deserves special attention since many probiotic strains within these genera are marketed for human consumption and, together with prebiotics, represent dietary strategies that shape either gut microbiota composition and/or its metabolic activity. The latter effects appear to be mediated not only at species but also at strain level. This points to the potential use of gut microbiota composition as predictive/therapeutic tool for NAFLD progression and severity. Notwithstanding this, microbiome-targeted interventions have shown promising but unsatisfactory clinical benefits. Microbiome-targeted therapies have shown significant immune or metabolic benefits for a minority of patients. The sharp influence of exogenous (i.e., diet and lifestyle, geography, prenatal nutrition) and endogenous (i.e., genes, ethnicity) factors on the transgenerational inheritance of heterogeneity in human gut microbiome composition still appears to represent a key step determining the clinical benefits for NAFLD patients. Overall, there is still a clinical need for contributions a) to understand the potential role of specific consortia of microbes; b) to foster interdisciplinary science establishing variations in gut microbiota composition and its genome within a ‘score of disease'; and c) to guide future therapeutic and management strategies for most patients.