Mi SciELO
Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en
SciELO
Similares en
SciELO -
 Similares en Google
Similares en Google
Compartir
Revista de Osteoporosis y Metabolismo Mineral
versión On-line ISSN 2173-2345versión impresa ISSN 1889-836X
Rev Osteoporos Metab Miner vol.10 no.4 Madrid nov./dic. 2018 Epub 03-Abr-2023
https://dx.doi.org/10.4321/s1889-836x2018000400003
Original Articles
Functional studies of DKK1 variants present in the general population
1Departamento de Genética, Microbiología y Estadística - Facultad de Biología - Universidad de Barcelona - Centro de Investigación Biomédica en Red de Enfermedades Raras (CIBERER) - Instituto de Biomedicina de la Universidad de Barcelona (IBUB) - Instituto de Investigación Sant Joan de Déu (IRSJD) - Barcelona (España)
2Departamento de Genética, Microbiología y Estadística - Facultad de Biología - Universidad de Barcelona - Barcelona (España)
3Unidad de Investigación en Fisiopatología Ósea y Articular (URFOA) - Instituto Hospital del Mar de Investigaciones Médicas (IMIM) - Parque de Salud Mar - Centro de Investigación Biomédica en Red de Fragilidad y Envejecimiento Saludable (CIBERFES) - Barcelona (España)
Objectives:
In recent decades, genes associated with bone mass and osteoporotic fracture risk have been identified, several of which belong to the Wnt pathway. In this project, the functionality of 7 missense mutations of the gene DKK1 –an inhibitor of the Wnt pathway– present in the general population was studied.
Materials and methods:
In vitro studies of the luciferase reporter gene were carried out to measure Wnt pathway activity in the presence or absence of wild-type or mutated DKK1, and western blot studies, to evaluate if the different mutations affect its synthesis and/or stability.
Results:
The DKK1 protein with the p.Ala41Thr variant shows lower pathway inhibitory activity compared to the wild-type protein. Significant differences were also observed between the experiments performed in the absence of DKK1 and those that include DKK1 with the p.Ala41Thr mutation. Western blots showed that the amount of protein was similar for all variants, both mutated and "wild-type, so the loss of p.Ala41Thr activity did not seem to be due to a lack of protein. The rest of the mutations did not show different behavior from that of the wild DKK1 protein.
Conclusions:
The missense variant p.Ala41Thr of the DKK1 protein, with a population frequency of 0.013%, shows a partial loss of its inhibitory function, which is not due to the lack of expression. This gene variant could lead to an increase in bone mineral density in those people in the general population who carry this mutation.
Key words DKK1; functional studies; missense variants; luciferase; Wnt pathway; High Bone Mass (HBM); osteoporosis
Introduction
The Wnt pathway’s role in regulating bone remodeling has been demonstrated in multiple studies. On the one hand, polymorphisms have been described in several genes of the Wnt pathway that show an association with bone mineral density (BMD) and the risk of fracture1-6. Rare or infrequent mutations have also been described in various genes of the Wnt pathway, which cause more rare bone phenotypes, such as osteoporosis-pseudoglioma (OPPG, OMIM 259770)7, autosomal recessive osteogenesis imperfecta of type XV (OMIM 615220)8, and osteosclerosis (OMIM 144750)9. The Wnt pathway begins with the formation of a heterotrimeric complex between the Frizzled receptor, the LRP5 co-receptor and the Wnt ligand. Once this complex is formed, β-catenin accumulates in the cytoplasm and translocates to the nucleus where it can activate the transcription of numerous target genes. In osteoblasts, the Wnt pathway has been shown to activate the transcription of genes that clearly contribute to bone formation10. In addition, this pathway is finely regulated by a series of extracellular inhibitors, including the protein sclerostin, encoded by the SOST gene, and the DKK1 protein, encoded by a gene with the same name. These two proteins perform their function, preventing the formation of the heterotrimeric complex. The proteins sclerostin and DKK1 thus form other heterotrimeric complexes, together with LRP5 and LRP4 (in the case of sclerostin)11,12 and together with LRP5 and Kremen (in the case of DKK1)13.
The transgenic over-expression of the DKK1 gene in osteoblasts produces a relative decrease in the number of osteoblasts compared to that of osteoclasts, thus producing a decrease in bone formation. Similarly, in mice, the homozygous deletion of the DKK1 gene is lethal, but deletion in heterozygosis presents a phenotype of bone overgrowth (high bone mass)14,15.
In the past decade, thanks to the direct effect on osteoblastogenesis inhibition and the indirect activation of osteoclastogenesis16, sclerostin and DKK1 have become interesting targets in osteoporosis treatment. Regarding DKK1, antibodies have been developed (BHQ880, DKN-01 and PF04840082), the first of which in the clinical trial phase in postmenopausal women with low bone mineral density (BMD)17-19.
In our group’s previous study20, DKK1 was sequenced to identify variants that could explain the high bone mass (HBM) phenotype , defined by a femoral + lumbar Z-score > 4, present in 15 women. In one of them, a missense mutation was found (p.Tyr74Phe) that co-segregated with the HBM phenotype in the family. In another study of DKK1 gene sequencing in postmenopausal women of the BARCOS cohort, we found another missense mutation (p.Arg120Leu) in another woman with HBM21. In addition to these mutations, in the general population there are other variants of change of direction in DKK1 (http://exac.broadinstitute.org/), whose effect in terms of bone mass has not been reported.
In the present work, we have conducted in vitro studies of the mutations p.Arg120Leu and p.Tyr74Phe, together with other missense mutations of DKK1 frequent in the general population (p.Met16Leu, p.Ala41Thr, p.Pro84Leu, p. Ala106Thr, p.Ser157Ile), to assess their possible involvement in bone phenotypes.
Material and method
Expression and mutagenesis vectors
The Wnt1-V5 mouse expression vectors, mesdc2, human wild-type LRP5, pRL-TK, PGL3-OT and DKK1-FLAG22, were courtesy of Dr. Wim van Hul (Antwerp, Belgium). Mutations p.Met16Leu, p.Ala41Tyr, p.Tyr74Phe, p.Pro84Leu, p.Ala106Thr, p.Arg120Leu, p.Ser157Ile were introduced into the DKK1-FLAG expression vector using the Quick Change Site-Directed Mutagenesis kit (Stratagene). The presence of mutations and the absence of errors were verified by Sanger sequencing.
Cell culture, production of conditioned medium and western blot
HEK293 cells, cultured with DMEM medium supplemented with FBS (10% V/V, Gibco, LifeTechnologies) and 1% streptomycin-penicillin (Gibco, LifeTechnologies) and maintained in incubators at 37°C at 5% in CO2. Were used. 3 x 105 cells were seeded per well in 6-well plates, 24 h before transfection. 2,000 ng/well of the mutated or wild-type DKK1-FLAG plasmids were transfected. The transfection was performed using Lipofectamine 2000 (Invitrogen) following the manufacturer's instructions. After 24 h, the medium was changed, reducing from 2 to 1 ml of DMEM, without FBS (Fetal Bovine Serum) or antibiotics. 48 hours after transfection, the supernatant of each condition was collected. The proteins of the conditioned medium were concentrated using Amicon Ultra filters (Millipore) and quantified by the BCA assay (Pierce). The proteins from the conditioned media (4.5 µg/lane) were separated by electrophoresis in a polyacrylamide gel with SDS (SDS-PAGE) and transferred to a nitrocellulose membrane. For the western blot analyses, Abcam ab109416 antibodies against DKK1 and ab2413 were used against the extracellular protein fibronectin, as a load control. The images were developed using a secondary antibody conjugated with peroxidase (Sigma-Aldrich). For each mutant conditioned medium was obtained in 2 different days and the analysis was carried out by western blot 2 times with these conditioned media.
Gene reporter assays
HEK293 cells were used, cultured as indicated in the previous section. 104 cells were seeded per well in 96-well plates, 24 h before transfection. Up to 5 plasmids were cotransfected in HEK293 cells: Mouse Wnt1-V5 (3.2 ng), mesdc2 (6.4 ng), human wild-type LRP5 (6.4 ng), pRL-TK (8 ng), and pGL3-OT (160 ng). In addition, depending on the experiment, the wild or mutated plasmid DKK1-FLAG (0.6 ng) was also co-transfected. If necessary, the empty vector pcDNA3 was used to equal the total amount of DNA from each experiment. The transfection was carried out using Lipofectamine 2000 (Invitrogen) following the manufacturer's instructions. 48 h after transfection, the cells were lysed and the luciferase activity of Photinus pyrali and Renilla reniformis was measured using a Glomax Multi+luminometer (Promega) following the instructions of the Dual-luciferase reporter assay (Promega). Each experiment included 5 replicates and was repeated independently in 3 separate experiments.
Statistical analysis
A one-way blocked ANOVA model was carried out for each mutant taking into account the test factor, the day as a blocking factor and the response variable the relationship between the activities of the luciferases (Photinus pyrali vs. Renilla reniformis). Blocking is a technique to deal with the nuisance factor and this can influence response. For each mutant protein, the test factor has the following levels: control (refers to the activity of the luciferase resulting from the endogenous Wnt pathway), the activator (luciferase activity produced by the Wnt pathway in the presence of Wnt and exogenous LRP5), the inhibitor (activity in the presence of the wild-type DKK1 inhibitor) and mutant (each of the mutant DKK1 proteins). The TukeyHSD test was used to carried out the post hoc test for multiple group comparisons. The ANOVA tests were done using the program R studio v.3.4.0, and values of p<0.05 were considered significant. All data were evaluated for normality, homogeneity of variance and detection of outliers.
Results
Expression, secretion and stability of mutated proteins
A western blot assay was performed to check if the DKK1 mutant proteins are correctly located in the extracellular space, using a culture of HEK293 cells, which express high amounts of wild-type DKK1 or mutated DKK1. The results show that, in all cases, the different mutated DKK1 proteins were detected in the extracellular space (Figure 1) and at levels equal to or higher than those of the wild-type protein.

Figura 1 Expression levels of the wild-type or mutated DKK1 protein analyzed by western blot. HEK293 cells were transfected with expression vectors of the different DKK1 variants indicated in each lane. The resulting conditioned media, properly concentrated, was used for this analysis. In each lane 4.5 ug of total protein was loaded. The extracellular protein fibronectin has been used as load control
Activity of mutated DKK1 proteins
To test the inhibitory activity of mutant DKK1 proteins on the Wnt pathway, we performed a reporter gene assay (luciferase), specific for this pathway (Figure 2).
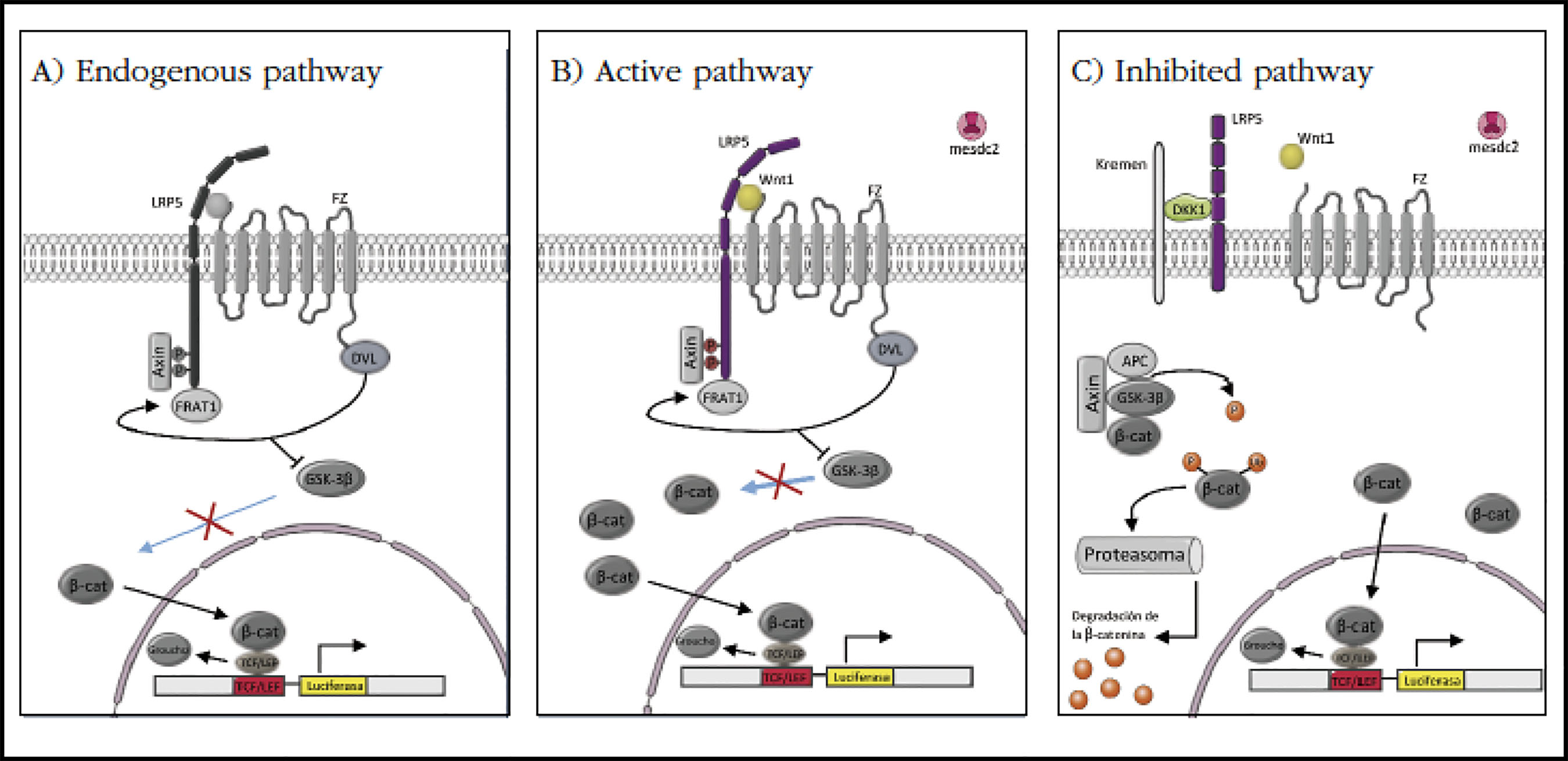
Figura 2 Reporter gene assay design. A) Endogenous condition: we transfected the pRLTK and pGL3OT plasmids. B) Active condition: we transfected plasmids pRLTK, pGL3OT, Wnt1, LRP5 and mesdc2. C) Inhibited condition: we transfected plasmids pRLTK, pGL3OT, Wnt1, LRP5, mesdc2 and DKK1-WT. In gray the endogenous elements of the HEK293 cells of the Wnt pathway, in color the transfected elements in each condition
The results of the endogenous condition in which the plasmids pRLTK and pGL3-OT have been co-transfected are shown in Figures 2A and 3 which represent the Wnt pathway activity in HEK293 cells.
In the active condition, in addition to pRLTK and pGL3-OT, the vectors expressing Wnt1 and LRP5 have been co-transfected. Wnt1 acts as a ligand activator of the pathway and LRP5 as a coreceptor, two essential elements for the pathway activation. In this condition (Figure 2B) the activity of luciferase has been increased 3 times, on average, compared to the endogenous pathway (Figure 3, activator).

Figura 3 Activity assay of the DKK1-p.Ala41Thr protein. A) Graph of interactions following the model, Yij = µ + day + test + day: test + ϵijk. The logarithm of the mean ratio of luciferase activities (Photinus pyrali/Renilla reniformis) with its confidence interval is shown on the Y axis. in the X axis the three days in which the experiment has been tested are shown. B) Tukey test of multiple comparisons between the different conditions tested. Significance levels are shown by means of the code: (****)<0.000001, (***)<0.0001, (**)<0.01, (*)<0.05
The inhibited pathway contained the same vectors as the active condition but in addition the vector expressing the protein DKK1-WT (wild-type protein) was co-transfected. In this condition, the Wnt pathway has been inhibited, by sequestering the LRP5 co-receptor (Figure 2C). In these experiments, luciferase activity has been increased 2.2fold over the endogenous condition and has been significantly lower than that of the activated pathway (Figure 3, inhibitor).
When the functionality of the mutants of DKK1 has been verified, the different vectors of the inhibited pathway have been co-transfected, but substituting that of DKK1-WT for those expressing the different mutated DKK1. For the mutant proteins DKK1p.Met16Leu, DKK1-p.Tyr74Phe, DKK1-p.Pro84Leu, DKK1-p.Ala106Thr, DKK1-p.Arg120Leu and DKK1p.Ser157Ile no significant differences were found in the inhibitory activity compared with the DKK1 WT protein (data not shown).
In contrast, in the presence of the mutant protein DKK1-p.Ala41Thr, luciferase activity has been observed which is significantly greater than that of the pathway inhibited by DKK1-WT (Figure 3), and in turn significantly lower than that of the active route.
Discussion
The DKK1 gene encodes a protein of the same name, which acts in the extracellular space as an inhibitor of the Wnt signaling pathway. Numerous studies have associated the Wnt pathway with bone formation, while blocking it with sclerostin or DKK1 has been associated with greater bone loss and risk of fracture. The search for gene variants that may explain Wnt pathway regulation in the general population may open a very relevant field of research in osteoporosis study. In this work, we have studied the inhibitory function of 7 mutant DKK1 proteins on the Wnt pathway and we have observed that the mutant protein DKK1p.Ala41Thr shows a partial loss of its inhibitory function, which is not due to the loss of its expression. The activity of the DKK1-p.A41T protein is reduced by approximately 50% compared to the protein DKK1-WT. The mutation, at amino acid 41, is not found in the LRP5 binding domain (amino acids 189-263), but it does affect the NAIKN motif (amino acids 40-44), which is crucial for binding to LRP5 and LRP6 proteins and it is conserved in all inhibitors of the Wnt pathway23.
According to the ExAC database, the population frequency of the variant p.Ala41Thr is 15 heterozygotes in 60,000 adult individuals free of serious diseases. Given our result of loss of inhibitory activity of this variant of DKK1, we could infer that the associated phenotype would be of a higher non-pathogenic bone density. From this frequency, we estimate that in Spain there are about 6,000 carriers of this variant in heterozygosis. On the other hand, there is a single reference to the mutation p.Ala41Thr that associates it with pathology, specifically the Chiari type I malformation (CMI)24. This disease is characterized by a defect in the development of the occipital bone and the posterior fossa (PF) and the consequent hernia of the cerebellar amygdala. It will be interesting to study the possible relationship between mutations in DKK1 and this disease, which, in many cases, is asymptomatic and undiagnosed.
No differences were found in the activity of the remaining mutant proteins and the DKK1 WT protein. These results coincide with the results found by Korvala et al.25 for the mutation p.Arg120Leu. These authors found this mutation in a patient with primary osteoporosis, a phenotype diametrically opposed to the phenotype of the HBM woman where we found the mutation. This same mutation is found in patients with Paget's disease (PDB)26 and its frequency in patients is twice that of controls, although the difference is not significant.
None of the seven mutations tested is in the domain that affects the Wnt signaling pathway (LRP5 binding domain: amino acids 189-263), and only p.Ala41Thr affects the NAIKN motif. This could be a reason why no differences in inhibitory activity have been observed in 6 of the 7 mutated DKK1. Alternatively, these DKK1 mutants would show differences in inhibitory activity lower than those that can be detected with the sensitivity of the reporter gene assay that has been used. A limitation of the study would be that the assay carried out involves the co-transfection of several vectors to have high values of luciferase activity, which gives it a high variability. Another limitation would be that the expected effect of these mutations is small, since they are variants present in the general population. This question can only be solved when there is a trial with a higher sensitivity.
In conclusion, in our study, DKK1 protein (p.Ala41Thr) shows a partial loss of its inhibitory function, which is not due to its lack of expression. This could lead to an increase in bone mineral density in people of the general population who carry this mutation.
REFERENCES
1 Freemantle N, Cooper C, Diez-Perez A, Gitlin M, Radcliffe H, Shepherd S, et al. Results of indirect and mixed treatment comparison of fracture efficacy for osteoporosis treatments: A meta-analysis. Osteoporos Int. 2013;24(1):209-17. [ Links ]
2 Peng J, Liu Y, Chen L, Peng K, Xu Z, Zhang D, et al. Bisphosphonates can prevent recurrent hip fracture and reduce the mortality in osteoporotic patient with hip fracture: A meta-analysis. Pakistan J Med Sci. 2016;32(2):499-504. [ Links ]
3 Kennel KA, Drake MT. Adverse effects of bisphosphonates: Implications for osteoporosis management. Mayo Clinic Proceedings. 2009;84(7):632-7. [ Links ]
4 Shane E, Burr D, Abrahamsen B, Adler RA, Brown TD, Cheung AM, et al. Atypical subtrochanteric and diaphyseal femoral fractures: second report of a task force of the American Society for Bone and Mineral Research. J Bone Miner Res. 2014;29(1):1-23. [ Links ]
5 Taormina DP, Marcano AI, Karia R, Egol KA, Tejwani NC. Symptomatic atypical femoral fractures are related to underlying hip geometry. Bone. 2014;63:1-6. [ Links ]
6 Sutton RAL, Mumm S, Coburn SP, Ericson KL, Whyte MP. “Atypical femoral fractures” during bisphosphonate exposure in adult hypophosphatasia. J Bone Miner Res. 2012;27(5):987-94. [ Links ]
7 Meier RPH, Lorenzini KI, Uebelhart B, Stern R, Peter RE, Rizzoli R. Atypical femoral fracture following bisphosphonate treatment in a woman with osteogenesis imperfectaa case report. Acta Orthop. 2012;83(5):548-50. [ Links ]
8 Alonso N, Soares DC, V McCloskey E, Summers GD, Ralston SH, Gregson CL. Atypical femoral fracture in osteoporosis pseudoglioma syndrome associated with two novel compound heterozygous mutations in LRP5. J Bone Miner Res. 2015;30(4):615-20. [ Links ]
9 Roca-Ayats N, Balcells S, Garcia-Giralt N, Falcó-Mascaró M, Martínez-Gil N, Abril JF, et al. GGPS1 mutation and atypical femoral fractures with bisphosphonates. N Engl J Med. 2017;376(18):1794-5. [ Links ]
10 Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754-60. [ Links ]
11 Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25(16):2078-9. [ Links ]
12 McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297-303. [ Links ]
13 Consortium T 1000 GP. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56. [ Links ]
14 Medina I, De Maria A, Bleda M, Salavert F, Alonso R, Gonzalez CY, et al. VARIANT: Command Line, Web service and Web interface for fast and accurate functional characterization of variants found by Next-Generation Sequencing. Nucleic Acids Res. 2012;40:W54-8. [ Links ]
15 Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073-81. [ Links ]
16 Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nature Methods. 2010;7(4):248-9. [ Links ]
17 Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 2005;15(8):1034-50. [ Links ]
18 Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44-57. [ Links ]
19 Boloc D, Castillo-Lara S, Marfany G, González-Duarte R, Abril JF. Distilling a Visual Network of Retinitis Pigmentosa Gene-Protein Interactions to Uncover New Disease Candidates. PLoS One. 2015;10(8):e0135307. [ Links ]
20 Chatr-Aryamontri A, Oughtred R, Boucher L, Rust J, Chang C, Kolas NK, et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 2017;45 (D1):D369-79; [ Links ]
21 Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. STRING v9.1: Protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41(D1):D808-15. [ Links ]
22 Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, et al. Circos: An information aesthetic for comparative genomics. Genome Res. 2009;19(9):1639-45. [ Links ]
23 Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: Archive for functional genomics data sets - Update. Nucleic Acids Res. 2013;41(D1):D991-5. [ Links ]
24 Yuen T, Stachnik A, Iqbal J, Sgobba M, Gupta Y, Lu P, et al. Bisphosphonates inactivate human EGFRs to exert antitumor actions. Proc Natl Acad Sci U S A. 2014;111(50):17989-94. [ Links ]
25 Brown JP, Morin S, Leslie W, Papaioannou A, Cheung AM, Davison KS, et al. Bisphosphonates for treatment of osteoporosis: expected benefits, potential harms, and drug holidays. Can Fam Physician. 2014;60(4):324-33. [ Links ]
26 Pérez-Núñez I, Pérez-Castrillón JL, Zarrabeitia MT, García-Ibarbia C, Martínez-Calvo L, Olmos JM, et al. Exon array analysis reveals genetic heterogeneity in atypical femoral fractures. A pilot study. Mol Cell Biochem. 2015;409:45-50. [ Links ]
27 Peris P, González-Roca E, Rodríguez-García SC, del Mar López-Cobo M, Monegal A, Guañabens N. Incidence of mutations in the ALPL, GGPS1 and CYP1A1 genes in patients with atypical femoral fractures. JBMR Plus. 2018; [Epub ahead of print]. [ Links ]
28 Zhou SF, Liu J-P, Chowbay B. Polymorphism of human cytochrome P450 enzymes and its clinical impact. Drug Metab Rev. 2009;41(2):89-295. [ Links ]
29 Napoli N, Villareal DT, Mumm S, Halstead L, Sheikh S, Cagaanan M, et al. Effect of CYP1A1 gene polymorphisms on estrogen metabolism and bone density. J Bone Miner Res. 2005;20(2):232-9. [ Links ]
30 Insalaco L, Gaudio F Di, Terrasi M, Amodeo V, Caruso S, Corsini LR, et al. Analysis of molecular mechanisms and anti-tumoural effects of zoledronic acid in breast cancer cells. J Cell Mol Med. 2012;16(9):2186-95. [ Links ]
31 Sims NA, Martin TJ. Coupling the activities of bone for mation and resorption: a multitude of signals within the basic multicellular unit. Bonekey Rep. 2014;3:481. [ Links ]
32 Perry JRB, Day F, Elks CE, Sulem P, Thompson DJ, Ferreira T, et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature. 2014;514(7520):92-7. [ Links ]
33 Mansouri MR, Schuster J, Badhai J, Stattin EL, Lösel R, Wehling M, et al. Alterations in the expression, structure and function of progesterone receptor membrane component-1 (PGRMC1) in premature ovarian failure. Hum Mol Genet. 2008;17(23):3776-83. [ Links ]
34 Xia WF, Tang FL, Xiong L, Xiong S, Jung JU, Lee DH, et al. Vps35 loss promotes hyperresorptive osteoclastogenesis and osteoporosis via sustained RANKL signaling. J Cell Biol. 2013;200(6):821-37. [ Links ]
35 Nguyen AM, Jacobs CR. Emerging role of primary cilia as mechanosensors in osteocytes. Bone. 2013;54(2):196-204. [ Links ]
36 Ducy P, Zhang R, Geoffroy V, Ridall AL, Karsenty G. Osf2/Cbfa1: A transcriptional activator of osteoblast differentiation. Cell. 1997;89(5):747-54. [ Links ]
37 Okada Y, Naka K, Kawamura K, Matsumoto T, Nakanishi I, Fujimoto N, et al. Localization of matrix metalloproteinase 9 (92-kilodalton gelatinase/type IV collagenase = gelatinase B) in osteoclasts: Implications for bone resorption. Lab Investig. 1995;72(3):311-22. [ Links ]
38 Nyman JS, Lynch CC, Perrien DS, Thiolloy S, O’Quinn EC, Patil CA, et al. Differential effects between the loss of MMP-2 and MMP-9 on structural and tissuelevel properties of bone. J Bone Miner Res. 2011;26(6):1252-60. [ Links ]
39 Pratap J, Javed A, Languino LR, van Wijnen AJ, Stein JL, Stein GS, et al. The Runx2 osteogenic transcription factor regulates matrix metalloproteinase 9 in bone metastatic cancer cells and controls cell invasion. Mol Cell Biol. 2005;25(19):8581-91. [ Links ]
Received: June 02, 2018; Accepted: September 01, 2018











 texto en
texto en 


