Mi SciELO
Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en
SciELO
Similares en
SciELO -
 Similares en Google
Similares en Google
Compartir
Revista de Osteoporosis y Metabolismo Mineral
versión On-line ISSN 2173-2345versión impresa ISSN 1889-836X
Rev Osteoporos Metab Miner vol.15 no.4 Madrid oct./dic. 2023 Epub 19-Feb-2024
https://dx.doi.org/10.20960/revosteoporosmetabminer.00029
SPECIAL ARTICLE
Polygenic risk scores (PRS) – A tool for disease prediction and personalized medicine
1Departamento de Medicina y Psiquiatría. Universidad de Cantabria. IDIVAL. Santander
2Servicio de Medicina Interna. Hospital Universitario Marqués de Valdecilla. Universidad de Cantabria. IDIVAL. CIBERER. Santander
Over the past decade, genomics and high-throughput sequencing have revolutionized our understanding of complex diseases. Polygenic risk scores (PRS) have emerged as a promising tool for predicting diseases and personalizing treatments. However, their implementation requires confirmation of real utility, which raises significant ethical and privacy challenges.
PRS are used to identify high-risk individuals and guide personalized treatments. Their potential is evident in diseases such as cancer or osteoporosis, where they improve risk stratification and enable the selection of more effective treatments. However, PRS have multiple limitations, including lack of individual accuracy, variability among different populations, and the inability to account for the impact of environmental factors. Clinical interpretation and ethical, legal, and social implications (ELSI) are highly relevant issues in this field.
In the future, PRS are expected to improve their predictive accuracy by combining clinical risk factors and adapting to populations of various ethnicities. Consequently, PRS are expected to play a central role in personalized medicine.
Keywords: Polygenic risk scores; Personalized medicine; Genome-wide association studies
INTRODUCTION
In the past decade, the advances made in genomics and high-throughput sequencing technology have transformed our understanding of the genetic basis of complex diseases. Polygenic risk scores (PRS) have emerged as an innovative tool that promises to revolutionize disease risk prediction and personalized medicine. These scores are based on the fundamental premise that many complex diseases, including heart disease (1), diabetes (2), cancer (3), or bone tissue disorders (4), result from the interaction of multiple genetic variants, each with a small effect individually. In this context, PRS allow for the calculation of individual genetic risk by adding information from hundreds, or even thousands, of genetic variants scattered throughout the genome (5).
The application of PRS in medical research and clinical care has prompted increasing interest due to its potential to identify individuals at higher risk of developing diseases, which, in turn, can guide early prevention and personalized management strategies. However, the implementation of PRS in the routine clinical practice has significant ethical, legal, and social implications (ELSI) and raises issues of privacy, result interpretation challenges, and the need to consider the clinical context and other risk factors (6).
This review aims to provide a non-exhaustive overview of PRS, from their theoretical foundation to their clinical application, and examine future perspectives and challenges faced in this field. Understanding this revolutionary tool is crucial in the context of disease genomics and has the potential to transform medical practice, providing new opportunities for the diagnosis, prevention, and individualized treatment of complex diseases.
GENERATION OF POLYGENIC RISK SCORES (PRS)
A polygenic risk score (PRS) represents an estimate of an individual's genetic burden related to a specific trait or disease. Its calculation is based on the sum of a subject's risk alleles, which are adjusted based on the effect size of these alleles, as derived from the results of genome-wide association studies (GWAS). The generation of PRS involves a series of fundamental methodological steps. An article published in Nature Protocols back in 2020 provides a detailed description of these steps, serving as a starting point and reference guide for researchers interested in conducting polygenic scoring analyses (7).
First, relevant genetic variants are selected from the GWAS results that have demonstrated a significant association with the disease of interest. Afterwards, these variants are then weighted according to their association strength, assigning weights that reflect their contribution to genetic risk. Subsequently, the “clumping” process groups the multiple variants, while considering linkage disequilibrium (LD) among them so that the SNPs retained are largely independent of each other. Additionally, the “thresholding” process involves applying a threshold to decide which variants are included in the PRS. This is done by considering the association strength of each variant, and if its P value or association statistic exceeds the specified threshold, it is included in the PRS. The PRS is calculated by adding the products of weighted variants by their alleles in the individual's genome. Eventually, its predictive capability is assessed using data from other independent groups (7,8).
In recent years, other methods have emerged to calculate PRS, such as the LASSO (Least Absolute Shrinkage and Selection Operator) method (9), which uses penalized regression to select informative SNPs by adding LD information, and Bayesian regression methods. Both have shown that they can achieve better performance than the “clumping + threshold” (C+T) method (10).
However, there is noticeable variability in the procedures for generating and validating PRS, making it challenging to compare and translate them into clinical care. The ClinGen Complex Diseases Working Group, in collaboration with the Polygenic Score Catalog (PGS), has updated the “genetic risk prediction information reporting” (GRIPS) to show the current state of the field. This document defines the minimum information required to interpret and evaluate PRS, especially in relation to subsequent clinical applications. Additionally, it emphasizes the importance of guaranteeing the availability and transparency of data, thus encouraging researchers to deposit and share PRS through PGS to facilitate replication in other studies (11).
CLINICAL APPLICATIONS
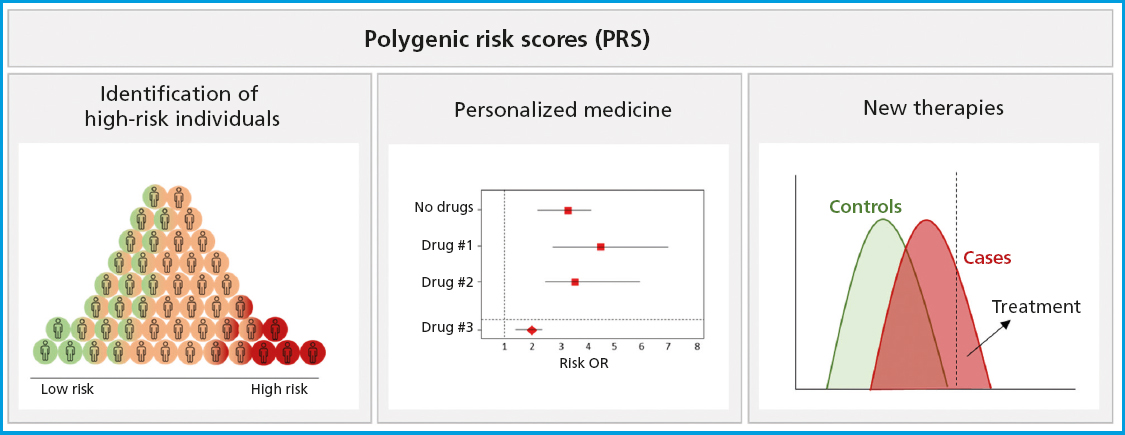
PRS have proven to be versatile tools in a wide range of clinical contexts and applications (Fig. 1). One of PRS prominent applications is the identification of high-risk individuals (11). By calculating a patient's PRS, individuals with significantly higher genetic risk of developing a disease compared to the overall population can be identified. This allows for more precise risk stratification and the possibility of early interventions, such as lifestyle changes, screening tests, or pharmacological prevention measures capable of reducing the incidence and severity of the disease.
Conventional non-genetic risk models typically add clinical and laboratory factors to identify high-risk individuals who are eligible for selective prevention strategies or prescription of medications to reduce the risk of disease. However, these factors fail to detect a significant number of individuals who will eventually develop the disease. For example, with cardiovascular risk calculators, almost 40 % of all individuals who will eventually suffer from heart diseases go unnoticed, especially if they are young individuals (12). Regarding breast cancer, the multiple factors used provide relatively weak predictions, identifying only a small proportion of individuals with long-term high risk (13).
In the field of metabolic bone diseases, the Fracture Risk Assessment Tool (FRAX) is a widely used tool to assess fracture risk. After initial analysis and validation in Spanish women without bone mineral density (BMD) analysis, the FRAX had a poor discrimination ability to predict major fractures but a good discrimination ability to predict hip fractures based on the area under the ROC curve. Nonetheless, the FRAX predictive ability is not much better than that from simple models based on age and BMD (14). More recently, using U.S. cut-off values, researchers confirmed that the FRAX performed better identifying patients who would not sustain major osteoporotic or hip fractures within the next 10 years compared to those who would. A considerable number of patients who developed major fractures were not identified in the initial assessment of the FRAX (15). In other words, based on these results, the FRAX specificity would be higher compared to its sensitivity.
Some studies have evaluated PRS along with the FRAX, as is the case with the PRS known as gSOS, which is related to fracture risk (16). The authors demonstrated that gSOS predicted the occurrence of a major osteoporotic, or hip fracture better than most traditional clinical risk factors, including previous fractures, use of corticosteroids, rheumatoid arthritis, and smoking, although always below the prediction level of BMD. They also demonstrated that adding gSOS to the FRAX improved the FRAX risk prediction ability, although still falling below the prediction level of FRAX + BMD (16). In a recent study, we analyzed the ability of 4 different PRS to predict osteoporosis among the Spanish population. We found that the osteoporosis group had a significantly higher genetic risk compared to the control group in 3 of these evaluated PRS. This suggests their potential utility in risk-based identification strategies based on a combination of clinical and genetic criteria (17).
Another important application is personalized medicine. PRS can guide the medical decision-making process adapted to a certain individual's genetic characteristics. PRS can help determine the optimal approach to treatment, selecting specific therapies that fit the patient's genetic profile and predicting what the patient's therapeutic response will be. In a study of therapies vs advanced breast cancer, the authors generated a final model with 13 single nucleotide polymorphisms (SNPs), which, when combined with the clinical covariates, showed predictive capability with a time-dependent area under the curve (AUC) of 0.81, compared to an AUC of 0.64 for the model with clinical covariates alone (18). This type of work demonstrates the potential of PRS in the field of pharmacogenomics to guide the selection of drugs and optimal dosages to maximize efficacy and minimize side effects. A systematic review aimed at obtaining information on the performance of PRS in predicting therapeutic outcomes identified a total of 89 articles that added pharmacogenetic variants into polygenic models. The authors could confirm that almost all studies found a significant association between their polygenic model and the outcome of the investigated medication (93 %). However, less than half (47 %) compared the performance of the polygenic model with clinical predictors, and only 40 % validated the model's predictions in an independent cohort (19). Manousaki et al. have explored the application of the previously described PRS-gSOS (16) as an independent risk factor for the occurrence of fractures in users of anti-osteoporotic drugs. They showed that patients with gSOS below average had increased adjusted odds (54 %) of sustaining major osteoporotic fractures and twice the adjusted odds of sustaining hip fractures compared to those with gSOS above average. Therefore, they demonstrated that PRS-gSOS is independently associated with incidental fractures among patients treated for osteoporosis (20).
All in all, although many association analyses have been conducted between polygenic risk and drug response in various fields such as anticoagulation, neuropsychiatric disorders, cancer, or various metabolic disorders (21), there are still many key considerations that should be made to improve and facilitate translation into the clinical setting of studies like these.
PRS are also used in the research of new therapies. By identifying individuals with a high genetic risk of a certain disease, PRS enable the selection of participants for prevention and treatment clinical trials. This facilitates the investigation of new therapeutic interventions, both pharmacological and non-pharmacological, with a specific focus on high-risk populations, thus accelerating the development of more effective treatments (22). Additionally, PRS contribute to the understanding of the genetic basis of these diseases, providing insights into the underlying biological pathways, which can lead to a better understanding of their pathogenesis and, ultimately, the development of more effective prevention and treatment (21,23).
LIMITATIONS
Despite their potential and utility in disease genetics and personalized medicine, PRS have several limitations that must be considered in their application and analysis. One of their main limitations is limited accuracy in individual prediction. Although PRS can provide an estimate of a person's genetic risk for a given disease, this estimation is relative and does not guarantee the occurrence of the disease. Genetic risk is only one of the various factors that impact the onset of complex diseases that does not taken into account environmental, lifestyle, or other risk factors that also play a crucial role. Therefore, they cannot be used exclusively to predict disease occurrence (24). The data analysis tools available today and the size of the cohorts available do not yet allow for in-depth analyses to consider the possible interactions between multiple genetic and environmental factors.
Additionally, the accuracy of PRS can vary depending on the population and ethnicity at stake. Most genome-wide association studies (GWAS) have been conducted in populations of European ancestry, which may limit the applicability of PRS in more diverse populations. In fact, several studies suggest that differences in the genetic structure of various populations can have an impact on the accuracy of PRS and may require specific adaptations for each population group (25,26).
Another important limitation is related to clinical interpretation. The information provided by PRS can be difficult to interpret for physicians and patients alike. PRS-based clinical decision-making requires a solid understanding of genetics and epidemiology, which may not be available in all clinical settings. Additionally, communicating PRS results to patients raises ethical and genetic counseling challenges (6,11).
ETHICAL AND PRIVACY CONSIDERATIONS
PRS raise important ethical and privacy issues. The personal genetic information contained in a PRS is sensitive and must be handled with caution. Protecting an individual's privacy and the safety of genetic data are critical issues when implementing PRS in the health care setting (27).
On the other hand, the implementation of PRS emphasizes the importance of properly communicating results to patients. Information on a person's genetic risk can be useful but can also generate anxiety and stress, especially if the interpretation of these results is not clear or if they are not accompanied by effective management recommendations. Appropriate genetic counseling and the communication of results in a judicious, understandable, and sensitive manner are essential to ensure that patients understand the meaning of their PRS and can make informed decisions on their health (11).
The privacy of genetic data is a critical concern in the use of PRS. Genetic information is highly sensitive and can reveal personal data such as ancestry and individual genetic characteristics. Protecting the privacy of genetic data is essential to prevent the misuse of this information, including genetic discrimination in areas such as employment or health insurance (28). Data safety is another concern. Genetic information must be stored securely to prevent unauthorized exposure or data theft. Implementing robust security practices and data encryption is essential to protect the privacy of all individuals and prevent vulnerabilities in the management of genetic data (29).
Finally, equity in access to genetic information is another relevant ethical issue. The availability of PRS can be biased toward those who have access to genetic sequencing services, thus leaving certain population groups behind. Ensuring equitable and fair access to PRS is a significant ethical challenge (30).
FUTURE PERSPECTIVES
The field of PRS continues to evolve and promises to play an increasingly relevant role in clinical genetics and precision medicine. Future perspectives in the field of PRS focus on improving the risks and limitations mentioned earlier. Greater predictive accuracy is required, which will improve as more evidence accumulates and more sophisticated models are developed. Adding whole-genome sequencing data and identifying rarer and lower-effect variants could significantly increase the predictive ability of PRS. Adaptation to diverse populations through the inclusion of data from different ethnic groups and populations will allow for a broader application of PRS on a global scale and improve their accuracy in non-European populations.
In the short term, and as can already be seen in various reports on different fields of medicine, including skeletal diseases, there are promising PRS, and it is likely that new PRS with greater sensitivity and specificity will emerge. These PRS, along with other risk factors (clinical, analytical, or imaging), could improve the stratification of patients at risk of bone fractures and design personalized preventive strategies. Something similar has already been implemented in some cancers, where strategies that combine gene panels and clinical factors are used to determine individual risk (31,32).
As PRS become more widespread, there will be more intense ethical and legal discussions on issues such as privacy, genetic discrimination, and equity in access. It will be essential to address these issues effectively and promote policies that protect individuals' rights.
Acknowledgements:
Álvaro del Real received support by the postdoctoral grant Margarita Salas in the University of Cantabria, Spain.
REFERENCES
1. O'sullivan JW, Raghavan S, Marquez-Luna C, Luzum JA, Damrauer SM, Ashley EA, et al. Polygenic Risk Scores for Cardiovascular Disease:A Scientific Statement from the American Heart Association. Circulation 2022;146(8):93-118. DOI:10.1161/CIR.0000000000001077 [ Links ]
2. Läll K, Mägi R, Morris A, Metspalu A, Fischer K. Personalized risk prediction for type 2 diabetes:the potential of genetic risk scores. Genet Med 2017;19(3):322-9. DOI:10.1038/GIM.2016.103 [ Links ]
3. Mavaddat N, Michailidou K, Dennis J, Lush M, Fachal L, Lee A, et al. Polygenic Risk Scores for Prediction of Breast Cancer and Breast Cancer Subtypes. Am J Hum Genet 2019;104(1):21-34. DOI:10.1016/J.AJHG.2018.11.002 [ Links ]
4. Forgetta V, Keller-Baruch J, Forest M, Durand A, Bhatnagar S, Kemp JP, et al. Development of a polygenic risk score to improve screening for fracture risk:A genetic risk prediction study. PLoS Med 2020;17(7):1003152. DOI:10.1371/JOURNAL.PMED.1003152 [ Links ]
5. Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet 2018;50(9):1219-24. DOI:10.1038/S41588-018-0183-Z [ Links ]
6. Lewis CM, Vassos E. Polygenic risk scores:from research tools to clinical instruments. Genome Med 2020;12(1). DOI:10.1186/S13073-020-00742-5 [ Links ]
7. Choi SW, Mak TSH, O'Reilly PF. Tutorial:a guide to performing polygenic risk score analyses. Nat Protoc. 2020;15(9):2759-72:44. 10.1038/S41596-020-0353-1. [ Links ]
8. Collister JA, Liu X, Clifton L. Calculating Polygenic Risk Scores (PRS) in UK Biobank:A Practical Guide for Epidemiologists. Front Genet 2022;13:818574. DOI:10.3389/FGENE.2022.818574 [ Links ]
9. Mak TSH, Porsch RM, Choi SW, Zhou X, Sham PC. Polygenic scores via penalized regression on summary statistics. Genet Epidemiol 2017;41(6):469-80. DOI:10.1002/GEPI.22050 [ Links ]
10. PrivéF, Arbel J, Vilhjálmsson BJ. LDpred2:better, faster, stronger. Bioinformatics 2021;36(22-23):5424-31. DOI:10.1093/BIOINFORMATICS/BTAA1029 [ Links ]
11. Wand H, Lambert SA, Tamburro C, Iacocca MA, O'Sullivan JW, Sillari C, et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 2021;591(7849):211-9. DOI:10.1038/S41586-021-03243-6 [ Links ]
12. Mars N, Koskela JT, Ripatti P, Kiiskinen TTJ, Havulinna AS, Lindbohm J V., al. Polygenic and clinical risk scores and their impact on age at onset and prediction of cardiometabolic diseases and common cancers. Nat Med 2020;26(4):549-57. DOI:10.1038/s41591-020-0800-0 [ Links ]
13. Brentnall AR, Cuzick J, Buist DSM, Bowles EJA. Long-term Accuracy of Breast Cancer Risk Assessment Combining Classic Risk Factors and Breast Density. JAMA Oncol 2018;4(9):180174. DOI:10.1001/JAMAONCOL.2018.0174 [ Links ]
14. Azagra R, Roca G, Encabo G, AguyéA, Zwart M, Güell S, et al. FRAX® tool, the WHO algorithm to predict osteoporotic fractures:the first analysis of its discriminative and predictive ability in the Spanish FRIDEX cohort. BMC Musculoskelet Disord 2012;13:204. DOI:10.1186/1471-2474-13-204 [ Links ]
15. Jiang X, Gruner M, Trémollieres F, Pluskiewicz W, Sornay-Rendu E, Adamczyk P, et al. Diagnostic accuracy of FRAX in predicting the 10-year risk of osteoporotic fractures using the USA treatment thresholds:A systematic review and meta-analysis. Bone 2017;99:20-5. DOI:10.1016/J.BONE.2017.02.008 [ Links ]
16. Lu T, Forgetta V, Keller-Baruch J, Nethander M, Bennett D, Forest M, et al. Improved prediction of fracture risk leveraging a genome-wide polygenic risk score. Genome Med 2021;13(1):16. DOI:10.1186/S13073-021-00838-6 [ Links ]
17. Real Á, Cruz R, Olmos Martínez JM, Hernández JL, Valero Díaz de la Madrid C, Riancho Moral JA. Polygenic risk of bone fractures in Spanish women with osteoporosis. Rev Osteoporos Metab Miner 2023;15(2):66-71. DOI:10.20960/REVOSTEOPOROSMETABMINER.00014 [ Links ]
18. Rashkin SR, Chua KC, Ho C, Mulkey F, Jiang C, Mushiroda T, et al. A Pharmacogenetic Prediction Model of Progression-Free Survival in Breast Cancer using Genome-Wide Genotyping Data from CALGB 40502 (Alliance). Clin Pharmacol Ther 2019;105(3):738-45. DOI:10.1002/CPT.1241 [ Links ]
19. Johnson D, Wilke MAP, Lyle SM, Kowalec K, Jorgensen A, Wright GEB, et al. A Systematic Review and Analysis of the Use of Polygenic Scores in Pharmacogenomics. Clin Pharmacol Ther 2022;111(4):919-30. DOI:10.1002/CPT.2520 [ Links ]
20. Manousaki D, Forgetta V, Keller-Baruch J, Zhao K, Greenwood CMT, Mooser V, et al. A Polygenic Risk Score as a Risk Factor for Medication-Associated Fractures. J Bone Miner Res 2020;35(10):1935-41. DOI:10.1002/JBMR.4104 [ Links ]
21. Cross B, Turner R, Pirmohamed M. Polygenic risk scores:An overview from bench to bedside for personalised medicine. Front Genet 2022;13. DOI:10.3389/FGENE.2022.1000667 [ Links ]
22. Zhou H, Mori S, Ishizaki T, Takahashi A, Matsuda K, Koretsune Y, et al. Genetic risk score based on the prevalence of vertebral fracture in Japanese women with osteoporosis. Bone Reports 2016;5:168-72. DOI:10.1016/J.BONR.2016.07.001 [ Links ]
23. Gibson G. On the utilization of polygenic risk scores for therapeutic targeting. PLoS Genet. 2019;15(4):1008060. DOI:10.1371/JOURNAL.PGEN.1008060 [ Links ]
24. Herzig AF, Clerget-Darpoux F, Génin E. The False Dawn of Polygenic Risk Scores for Human Disease Prediction. J Pers Med 2022;12(8):1266. DOI:10.3390/JPM12081266 [ Links ]
25. Roberts MC, Khoury MJ, Mensah GA. Perspective:The Clinical Use of Polygenic Risk Scores:Race, Ethnicity, and Health Disparities. Ethn Dis 2019;29(3):513-6. DOI:10.18865/ED.29.3.513 [ Links ]
26. Evans DG, van Veen EM, Byers H, Roberts E, Howell A, Howell SJ, et al. The importance of ethnicity:Are breast cancer polygenic risk scores ready for women who are not of White European origin?Int J cancer. 2022;150(1):73-9. DOI:10.1002/IJC.33782 [ Links ]
27. Adeyemo A, Balaconis MK, Darnes DR, Fatumo S, Granados Moreno P, Hodonsky CJ, et al. Responsible use of polygenic risk scores in the clinic:potential benefits, risks and gaps. Nat Med 2021;27(11):1876-84. DOI:10.1038/S41591-021-01549-6 [ Links ]
28. Lewis ACF, Green RC. Polygenic risk scores in the clinic:new perspectives needed on familiar ethical issues. Genome Med 2021;13(1):14. DOI:10.1186/S13073-021-00829-7 [ Links ]
29. Wan Z, Hazel JW, Clayton EW, Vorobeychik Y, Kantarcioglu M, Malin BA. Sociotechnical safeguards for genomic data privacy. Nat Rev Genet 2022;23(7):429-45. DOI:10.1038/S41576-022-00455-Y [ Links ]
30. Khoury MJ, Bowen S, Dotson WD, Drzymalla E, Green RF, Goldstein R, et al. Health equity in the implementation of genomics and precision medicine:A public health imperative. Genet Med 2022;24(8):1630-9. DOI:10.1016/J.GIM.2022.04.009 [ Links ]
31. Evans DGR, van Veen EM, Harkness EF, Brentnall AR, Astley SM, Byers H, et al. Breast cancer risk stratification in women of screening age:Incremental effects of adding mammographic density, polygenic risk, and a gene panel. Genet Med 2022;24(7):1485-94. DOI:10.1016/J.GIM.2022.03.009 [ Links ]
32. Roberts E, Howell S, Evans DG. Polygenic risk scores and breast cancer risk prediction. Breast 2023;67:71-7. DOI:10.1016/J.BREAST.2023.01.003 [ Links ]
Received: November 08, 2023; Accepted: December 09, 2023











 texto en
texto en